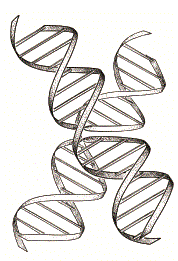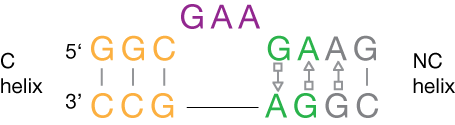Nucleic Acid Structure Research Group
University of Dundee
Research
Skip to:
DNA junctions
RNA structure and activity
Development of methods for the study of branched nucleic acids
Structure, Folding and Activity of Branched Nucleic Acids
Helical branchpoints are important elements in both DNA and RNA. In DNA they constitute central intermediates in recombination and repair processes, while in RNA they are important three-dimensional architectural elements.
There is a universal nomenclature for nucleic acid junctions, adopted by IUB (Lilley et al., 1995).
DNA structures
DNA junctions
DNA branchpoints are created during a number of important processes in the cell, including repair, recombination and replication (Lilley, 2000). Genetic recombination is a process by which similar DNA molecules become transiently broken and then re-joined in a new combination. It provides genetic diversity, the raw material of evolution, and an important method by which cells can repair their genetic material against accidental breakage. The central intermediate of this process is a four-way junction (or Holliday junction), a structure in which four double-helical DNA molecules are connected by mutually exchanging component strands (Holliday, 1964).
Electron micrograph of a four-way (Holliday) DNA junction. Original micrograph provided by Dr H Potter (Potter and Dressler, 1978).
We determined the structure of this species in 1988, using electrophoretic and fluorescence approaches (Duckett et al., 1988; Murchie et al., 1989).
The stacked X-structure of the DNA junction
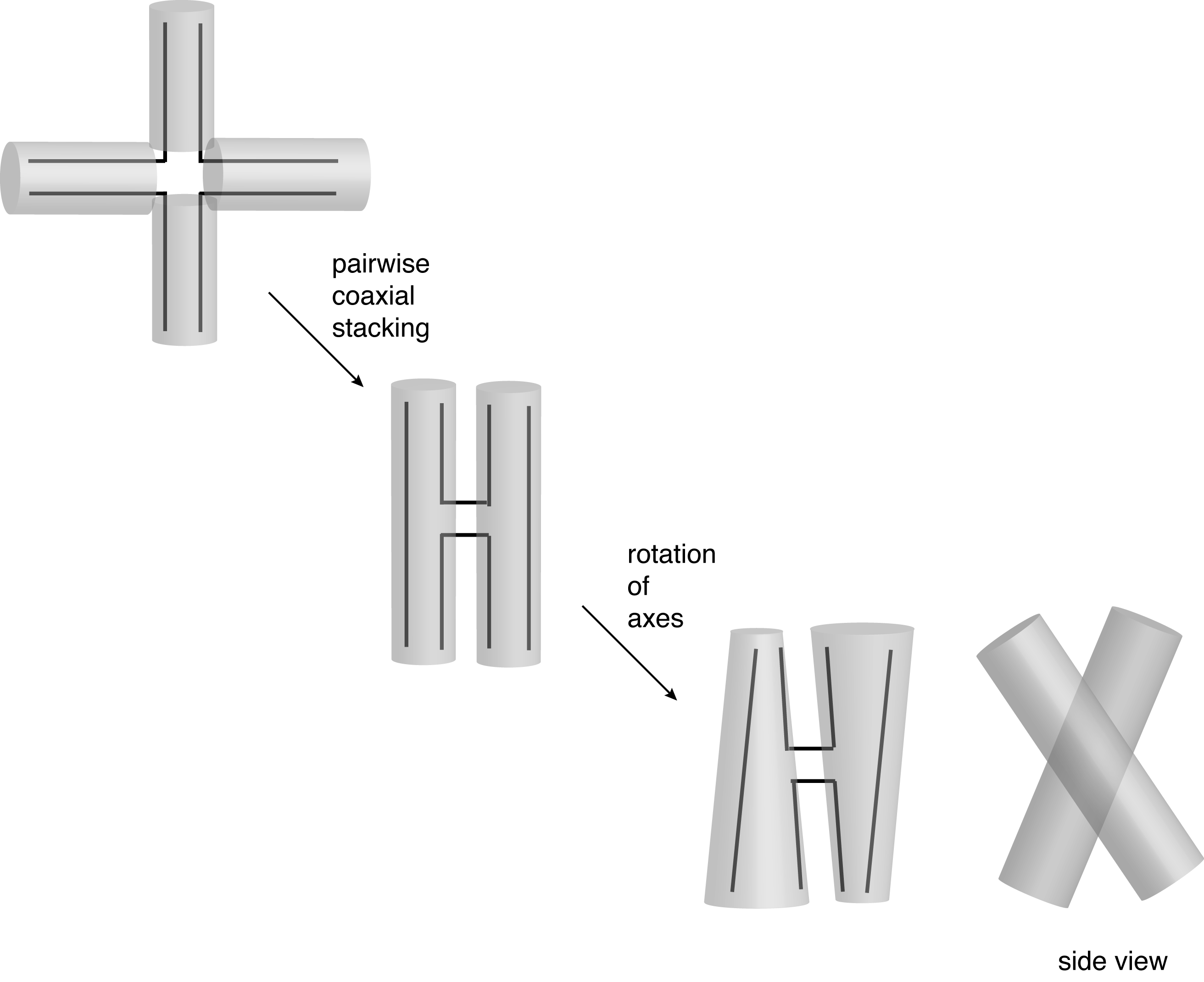
In the presence of metal ions, the junction adopts the stacked X-structure by the pairwise, coaxial stacking of helices. This is a right-handed, antiparallel structure. The structure can be considered to be formed by coaxial stacking of helices together with a rotation of the axes by 40-60° in a right-handed sense.
|
|
Folding of the four-way DNA junction into the stacked X-structure. Left, a conceptual path to the folded structure by pairwise coaxial stacking, followed by rotation of the axes. Right, a ribbon depiction of the stacked X-structure.
When the original structure of the stacked X-structure was determined the data new indicated that it must be antiparallel, in contrast to earlier assumptions and the virtually universal parallel depiction in text books. More recent single-molecule studies have shown that the probability of parallel forms of the junction with a lifetime greater than 1 ms is less than 2 x 10-5 (Joo et al., 2004).
The structure was confirmed in virtually every detail by X-ray crystallography more than a decade after our original determination of its structure (Eichman et al., 2000; Ortiz-Lombardía et al., 1999; Thorpe et al., 2003).
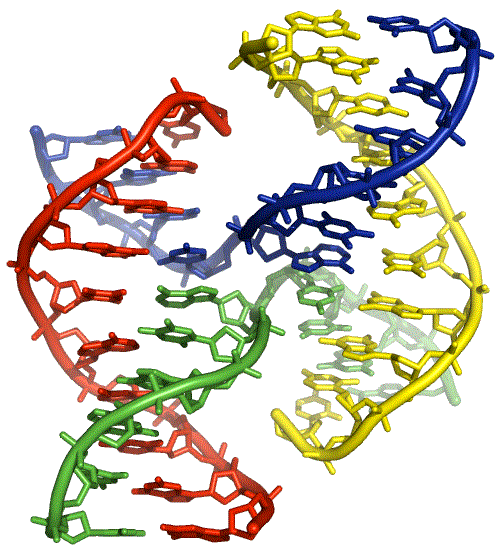
A crystal structure of a four-way DNA from Shing Ho and colleagues (Eichman, 2000). The junction adopts the right-handed stacked X-structure. The red and yellow strands are continuous strands, while the blue and green strands exchange at the centre. The continuous strands run antiparallel. The view shown is the major groove side of the junction.
Electrostatics, metal ions and the stacked X-structure
Nucleic acids are highly charged polyelectrolyte molecules, and like most such species their structure depends strongly on the presence or absence of metal ions such as magnesium. Simple electrostatic calculations show that there should be a regions of very high electrostatic potential at the center of the stacked X-structure, and metal ions will be required to neutralize the charge repulsion. In the absence of this, the repulsive energy dominates and the extended open structure will be more stable. The overall shape of the low-salt form would be expected to be square as this provides the maximal extension around the centre, and would therefore probably minimize electrostatic repulsion. But this would not necessarily be planar; given that the two sides of the extended structure are inequivalent (there are major and minor groove faces) this is indeed unlikely, and the structure is probably to some degree pyramidal.
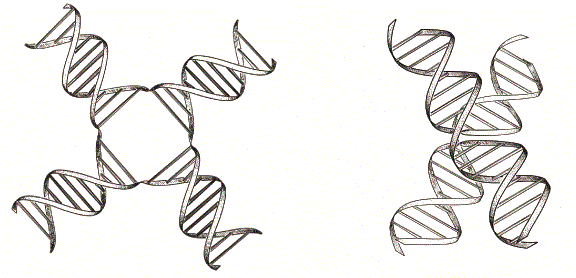
Two forms of the four-way DNA junction. In the absence of ions, electrostatic repulsion results in a square structure with an open center (left), in which the four helical arms point to the four corners of a square. On lowering the electrostatic interactions between charged phosphate groups by addition of metal ions, the junction can undergo a folding process by the pairwise coaxial of helical arms to form the stacked X-structure (right).
Steady-state ensemble FRET analysis of the ion-induced folding of the four-way junction indicates that the process is well described by a simple two-state model (Clegg et al., 1992; Fogg et al., 2001). The experimental data can be fitted with a Kaapp for ion divalent metal ion binding of ~105 M-1, and a Hill coefficient of n = 1. This is consistent with a model in which folding is induced by the non-cooperative binding of one metal ion. However this is unlikely to be site bound as an inner-sphere complex for two reasons. First, folding can be induced using monovalent metal ions alone, and second, no metal ion bound at the point of strand exchange has been identified by crystallography (Eichman et al., 2000; Thorpe et al., 2003).

Ion-induced folding of the four-way DNA junction. FRET titration with Mg2+ ions. A junction was labelled with donor and acceptor fluorophores attached to the termini of two arms that define one of the acute angles of the folded junction. The efficiency of fluorescence resonance energy transfer (EFRET) was measured as a function of Mg2+ ion concentration. The data were fitted to a simple two-state model for folding induced by ion binding, such that the proportion of folded junction (&alpha) is given by:
Kaapp [Mg2+] n / ( 1 + Kaapp.[Mg2+]n), where Kaapp is the apparent association constant for Mg2+ ion binding and n is the Hill coefficient. The simplest interpretation is that folding is induced by the non-cooperative binding of a single Mg2+ ion.
Nevertheless, it is likely that there is a high occupancy of rapidly exchanging metal ions at the center of the junction, as outer-sphere complexes. Uranyl-induced photocleavage experiments indicated the presence of an ion binding site near the point of strand exchange in the folded junction (Mřllegaard et al., 1994). The two phosphates that define the point of strand exchange on the exchanging strands are only separated by 6.5 Ĺ, and there is a box of six phosphates comprising these two plus those 5'and 3' to them on each strand.
| |
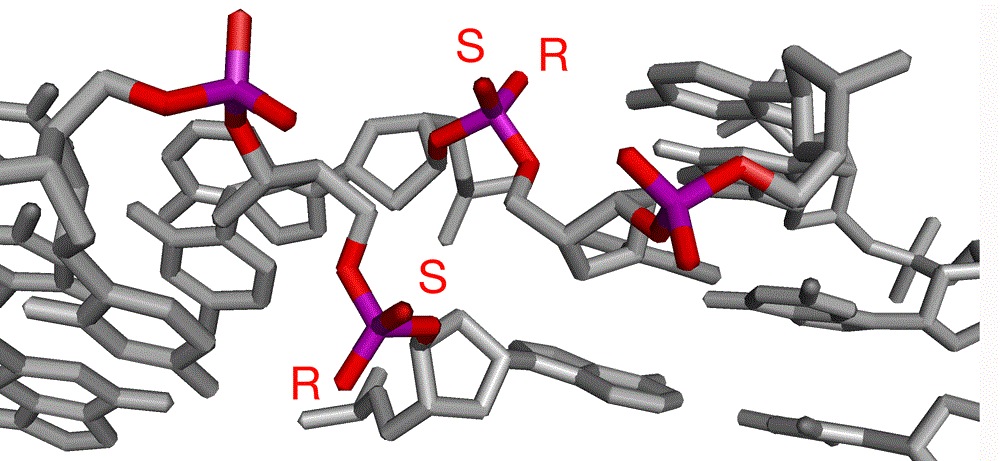
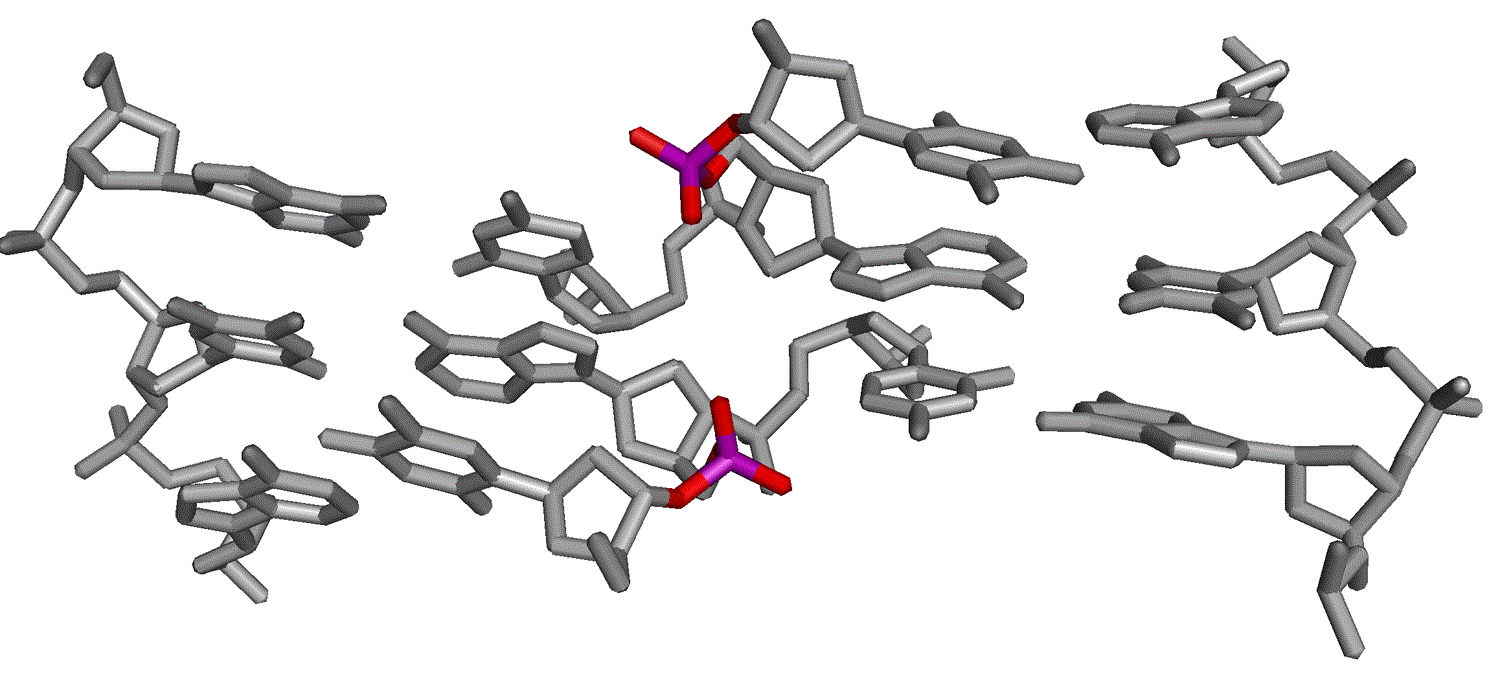
Phosphate groups at the center of the four-way DNA junction. Left, minor groove side showing central and 3'-phosphates. The stereoscopic identity of the non bridging oxygen atoms of the central phosphates have been labeled. Right, the major groove side, showing the 5'-phosphates.
Replacement of any of these phosphates by the electrically-neutral methyl phosphonate groups increases the stability of the folded form of the junction at lower ionic concentrations (Liu et al., 2004). However, these effects can be rather more subtle. When methyl phosphonate groups were substituted for the central phosphate on one of the continuous strands of a junction (in its major conformer) it was found that two stacking conformers co-existed in solution without interconversion (Liu et al., 2005). Further analysis showed that these correspond to the alternative diastereomers of the substituted phosphate group. It is therefore probable that the location of the methyl group determines the relative stability of the alternative stacking conformers due to differential interactions with metal ions at the point of strand exchange. Substitution of the pro-R oxygen atom stabilizes the folded junction, and therefore represents the simple electrostatic effect. But substitution of the pro-S oxygen atom destabilizes the stacked X-structure, probably arising from steric interference with ionic interactions.
The DNA junction as a dynamic structure
There are two ways that a given junction can be folded into the stacked X-structure, depending on the choice of stacking partner. The two structures are stereochemically equivalent, but will usually differ in terms of the stacking interactions across the junction.
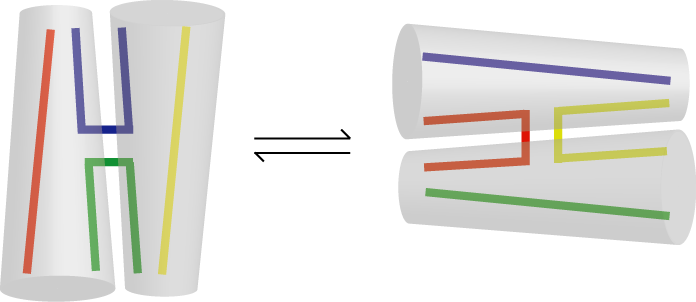
Alternative stacking conformers of the four-way DNA junction. Note that the conformational transition exchanges continuous and exchanging strands.
Thus they will not be energetically equivalent, and so most junctions exhibit a distinct conformational bias towards one conformer. The bias has never been found to be total however, and a significant population of the minor conformer has been found in all junctions studied. Moreover, at least one sequence (termed junction 7) was identified where the populations of the two conformers were very close to being equal (Grainger et al., 1998).
The existence of two populations of stacking conformers suggested that the two forms might interconvert. There is no way to synchronize conformer exchange, and it was necessary to use single-molecule methods to demonstrate this directly. FRET between fluorophores terminally-attached to different arms of single junction molecules revealed the exchange occurring between junction conformers (McKinney et al., 2003), occurring at a surprisingly slow rate in the presence of 50 mM Mg2+ ions.
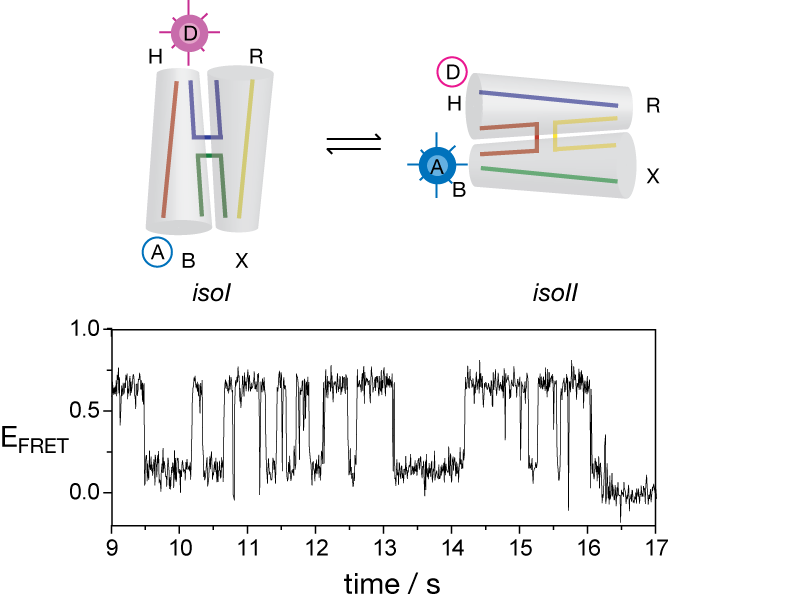
Conformer exchange in a four-way junction studied by single-molecule FRET. A record of FRET efficiency (EFRET) as a function of time for a single junction molecule shows repeated interconversion between the low efficiency (EFRET = 0.1) and the high efficiency (EFRET = 0.6) states (McKinney et al., 2003).
These experiments in collaboration with Taekjip Ha further showed that the rate of conformer exchange became significantly faster as the ionic strength of the solution was lowered (Joo et al., 2004). This indicates that the transition state is destabilized by metal ions, suggesting that it resembles the open form of the junction that exists in the absence of metal ions. This makes structural sense, since it is hard to imagine how the required exchange of stacking partners could occur without substantial opening of the junction. The lifetime of the open species was estimated to be < 1 ms. Conformer exchange has been studied by the application of stretching force (Hohng et al., 2007). By measuring the gradient of the rate of conformer exchange as a function of applied force it was possible to determine the position of the transition state along the reaction coordinate. Using relatively low forces (< 4 pN) it was determined that the open structure is a shallow intermediate, flanked by transition states on either side. The application of stretching force tilts the entire energy landscape, thereby altering the prevailing rate determining step.
In homologous junctions there is no fixed position for the branchpoint, which can therefore change position due to the exchange of basepairing, termed branch migration. Branchpoints can migrate over long distances, and the process occurs as a step-wise random walk. A single step of branch migration requires the breakage of two basepairs at the point of strand exchange located on diametrically opposite arms, rotation, and pairing to form the terminal basepairs on the other two arms. This would be expected to require substantial opening of the junction, much as envisaged for conformer exchange. The process was studied in single junction molecules that could undergo one step of branch migration (McKinney et al., 2005). Individual steps of branch migration could be observed, together with the conformer exchange occurring in each form of the junction. It was concluded that conformer exchange and branch migration share a common intermediate that is likely to resemble the open state deduced for free junctions in the absence of divalent metal ions.
Recognition of DNA junctions by proteins
The structure of four-way DNA junctions is recognized by a number of proteins, especially the group of nucleases that introduce specific pairs of cleavages in order to resolve the junction into nicked duplex species (Déclais and Lilley, 2008).
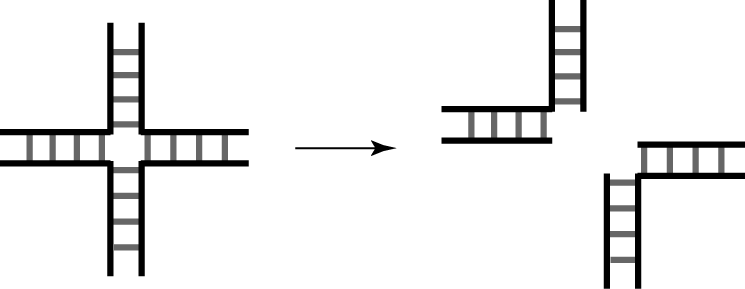
Junction-resolving enzymes are selective for the structure of four-way DNA junctions, introducing paired cleavages in order to resolve them into two DNA species that may be ligated to reform duplex structures.
Junction-resolving enzymes have been isolated from many organisms, from bacteria and their phage, yeast, archaea, through to mammalian cells and their viruses (Lilley and White, 2001; West, 2003). The enzymes bind four-way DNA junctions as dimers, with affinities close to Kd = 1 nM. Binding is highly structure selective; the complex of a dimer of a given junction-resolving enzyme and a four-way DNA junction is normally not disrupted by a 1000-fold excess of duplex DNA of the same sequence. Yet despite recognizing junction structure, all the junction-resolving enzymes distort the very structure that they recognize. Many substantially open the structure of the junction to different degrees - the most extreme case is Cce1, where the junction adopts an open-square structure like that of the free junction in the absence of metal ions (White and Lilley, 1997). In addition, 2-aminopurine fluorescence experiments suggested that the binding of Cce1 results in a significant disruption of base pairing adjacent to the point of strand exchange in addition to the global change (Déclais and Lilley, 2000).
Tabulation of junction-resolving enzymes. Most fall into one of two groups of proteins, the integrase family and the nuclease family (Lilley and White, 2000). Phage T4 endonuclease VII and DPL12 cannot be classified with these enzymes.
The structural distortion of the DNA junctions induced by enzyme binding may be related to the their function, helping to assure a well-ordered resolution of the junction. Detailed kinetic analysis of the cleavage of cruciform structures revealed that there is an acceleration of the second strand cleavage (by a factor of 100 in the case of RuvC), so that it immediately follows cleavage of the first strand (Fogg and Lilley, 2001; Fogg et al., 2000; Giraud-Panis and Lilley, 1997). We suggested that the intact junction is strained in the complex containing the intact junction, and the first cleavage event is followed by a relaxation that accelerates the second cleavage.
Crystal structures have been determined for most of the proteins in isolation. (Ariyoshi et al., 1994; Bond et al., 2001; Ceschini et al., 2001; Hadden et al., 2001; Kelly et al., 2007) However, the fundamental question of the origins of structural recognition required the determination of the structures of complexes of the resolving enzymes with DNA junctions. This has now been achieved for the phage enzymes T7 endonuclease I (Hadden et al., 2007; McGregor et al., 2005; Middleton et al., 2004; Nishino et al., 2001; Raaijmakers et al., 1999) and T4 endonuclease VII (by Wei Yang and colleagues (Biertümpfel et al., 2007). And it turns out that these two proteins recognize the structure of the junction in rather different ways.
T4 endonuclease VII (Biertümpfel et al., 2007) distorts the junction into an almost planar open X-shape. The DNA is bound via its minor groove face to a fairly flat surface of the enzyme, making contacts with all four helical arms. By contrast, when a junction is bound by T7 endonuclease I (Hadden et al., 2007) the helical arms remain coaxial (though the axes are rotated by 130°), conferring a much more three-dimensional aspect to its shape.
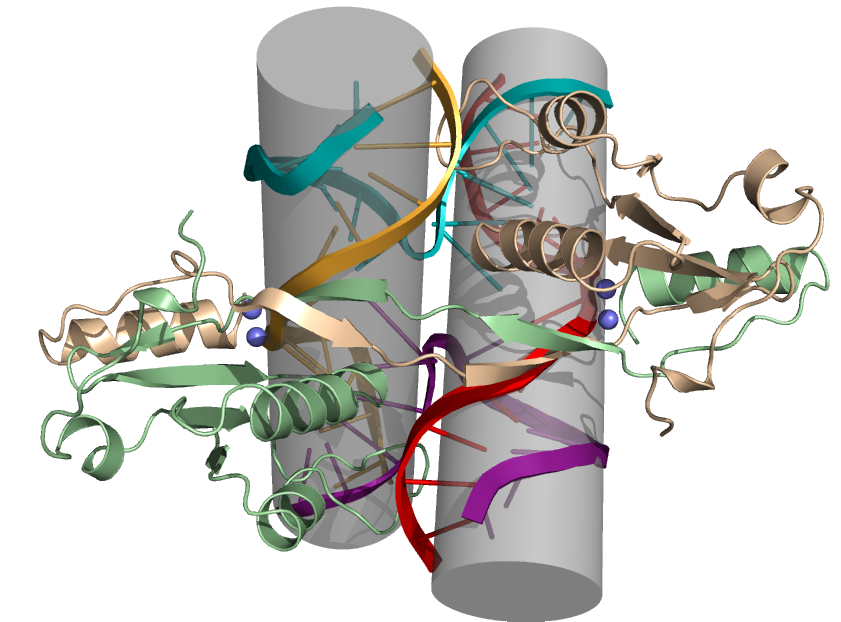
Crystal structure of a complex of a four-way junction with the junction-resolving enzyme endonuclease I of bacteriophage T7 (collaboration with Simon Phillips). The protein is shown in cartoon form, with the two metal ions within each active site shown by the blue spheres.
The protein dimer presents two 30 Ĺ-long hemi-cylindrical clefts that are mutually perpendicular.

Crystal structure of a complex of a four-way junction with T7 endonuclease I. The protein is shown spacefilling to highlight the formation of the long clefts in which the stacked helical arms of the junction are bound,
The DNA arms are bound along the length of these electropositive channels, and thus the enzyme is selective for a DNA structure that can adopt the near-perpendicular geometry of the junction (Déclais et al., 2006; Hadden et al., 2007). The active sites of the enzyme (Déclais et al., 2001) are also located in the grooves, where two metal ions (Hadden et al., 2002) provide the hydrolytic water molecule for nucleophilic attack on the scissile phosphate groups of the continuous strands in a mechanism (Liu et al., 2006) that is closely similar to that of many restriction enzymes.
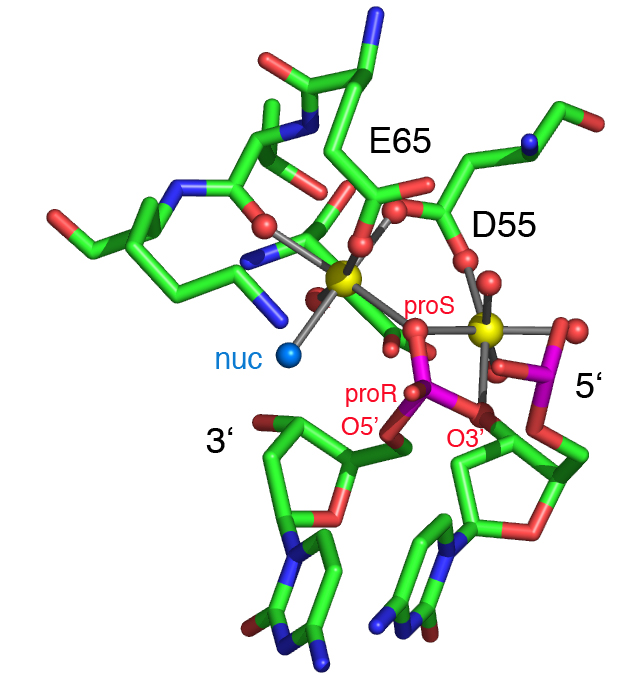
A model of the active site of T7 endonuclease I with a dinucleotide bound (Liu et al., 2006). The two metal ions are shown as yellow spheres, with associated water of hydration. The water molecule (nuc, highlighted cyan) coordinated to metal ion 1 is well positioned for in-line attack on the scissile phosphorus opposite the 3' oxygen leaving group. The proS oxygen atom is coordinated to both metal ions (replacing a water molecule that bridges the ions in the free protein), while the proR does not interact with either ion. The coordination of the two metal ions in this model is completely equivalent to that observed in the BglI-DNA complex (Newman et al., 1998).
Eukaryotic junction-resolving enzymes
Nuclear junction-resolving enzymes from eukaryotes have been elusive, but significant progress in their identification and isolation has been made recently. Human Gen-1 and its yeast ortholog was identified by Steve West and colleagues (Ip et al., 2008). It appears to have the properties expected of a bona fide junction-resolving enzyme. With our colleague Anton Gartner we have identified the corresponding enzyme in C. elegans.
A second human junction-resolving enzyme has recently come to light. With our colleague John Rouse, we (and several other laboratories) have shown that the complex of SLX1 and SLX4 has junction-resolving activity (Munoz et al., 2009).
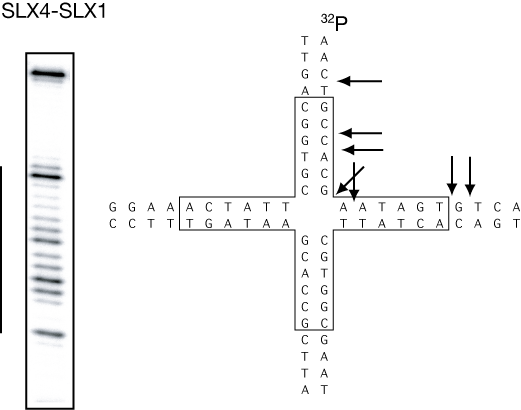
Cleavage of a DNA junction by the complex of SLX1-SLX4. The junction can branch migrate over the central region (boxed), and the enzyme makes a number of cleavages within and just outside of this region (shown by the line to the left of the gel).
In addition, we and others have begun to study nucleases from eukaryotes that are targeted to three-way junctions containing nicks (possibly related to replication intermediates) and related flap structures, such as the FAN1 protein (MacKay et al., 2010) .
RNA structure and activity
RNA junctions
Helical junctions are common elements in the secondary structure of RNA. They have a key role in the folding of small functional RNA molecules such as some of the nucleolytic ribozymes and the HCV IRES element, and are important in larger RNA species, organising the tertiary structure of the molecule. Interestingly, if we take the nucleolytic ribozymes as a representative collection of small autonomously-folding RNA species we find they divide into two groups. The structure of three (the hammerhead, hairpin and VS ribozymes) are based around helical junctions, while the remaining two (the hepatitis delta virus and glmS riboswitch ribozymes) are instead based on complex, nested pseudoknots. For these molecules at least, these two ways to solve the RNA folding appear to be mutually exclusive. In larger RNA species we can dissect the folding problem into the interactions between helices. These will be of two kinds, tertiary interactions, such as between loops and their receptors, and junctions that connect the helical segments. Our philosophy has therefore been to study the conformational properties of RNA junctions in depth.
Perfectly-paired (4H) junctions exist in a number of significant RNA species, such as the hairpin ribozyme and U1 snRNA (Duckett et al., 1995). Like their DNA equivalents, they undergo pairwise coaxial stacking of helices, though the structures are rather more polymorphic. Single-molecule FRET has shown (Hohng et al., 2004) that they undergo exchange between alternative stacking conformers, but also (in marked contrast to DNA junctions) between parallel and antiparallel forms.
Perfectly-paired junctions are actually rather rare in natural RNA species, and most have one or more formally-unpaired nucleotides linking the helices. Unsurprisingly, the structural and dynamic properties cane be altered by the presence of the additional nucleotides, as has been shown to be the case for the 2HS22HS1 four-way junction found in the HCV IRES element (Melcher et al., 2003). In contrast to 4H RNA junctions, the IRES junction was found to lose coaxial stacking and adopt an extended structure in the absence of added metal ions. On addition of divalent metal ions the junction folded by pairwise coaxial stacking of arms, placing the extra nucleotides onto the exchanging strands. Time-resolved FRET studies indicated the existence of a rapid exchange between approximately equal populations of parallel and antiparallel conformations.
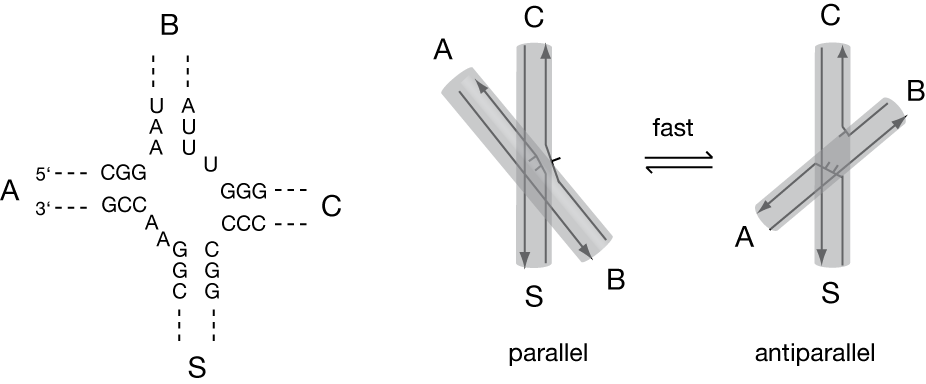
The four-way RNA junction of the HCV IRES (Melcher et al., 2003). The central sequence of the junction is shown (left). The junction undergoes a rapid exchange between parallel and antiparallel conformations (right).
Three-way junctions are the most common branchpoints found in natural RNA molecules. They are seldom found as simple 3H junctions, but occur with of varying degrees of complexity. The presence of additional nucleotides between the helical sections increases conformational flexibility. Analysis of three-way junctions in crystallographic structures reveals that they generally fold by the coaxial stacking of two of the three arms. A good example of this is the three-way junction of the hepatitis C-virus internal ribosome entry site (Ouellet et al., 2010). Three-way junction conformation has been analysed by Lescoute and Westhof (Lescoute and Westhof, 2006).
The Varkud Satellite ribozyme provides a good example of the importance of three-way junctions in setting up the basic framework of a functional RNA molecule. The secondary structure (Beattie et al., 1995) suggests that the basic core of the ribozyme is organized by two three-way junctions. These direct the trajectory of five helices (numbered II through VI) into an H-shape. Helices III, IV and V create a HS1HS5HS2 junction, while helices II, III and VI form a 2HS5HS2 junction. Both junctions undergo two-state folding induced by the non-cooperative binding of divalent metal ions (Lafontaine et al., 2001a; Lafontaine et al., 2002). This information was later incorporated into a general model for the structure of the VS ribozyme (Lipfert et al., 2008).
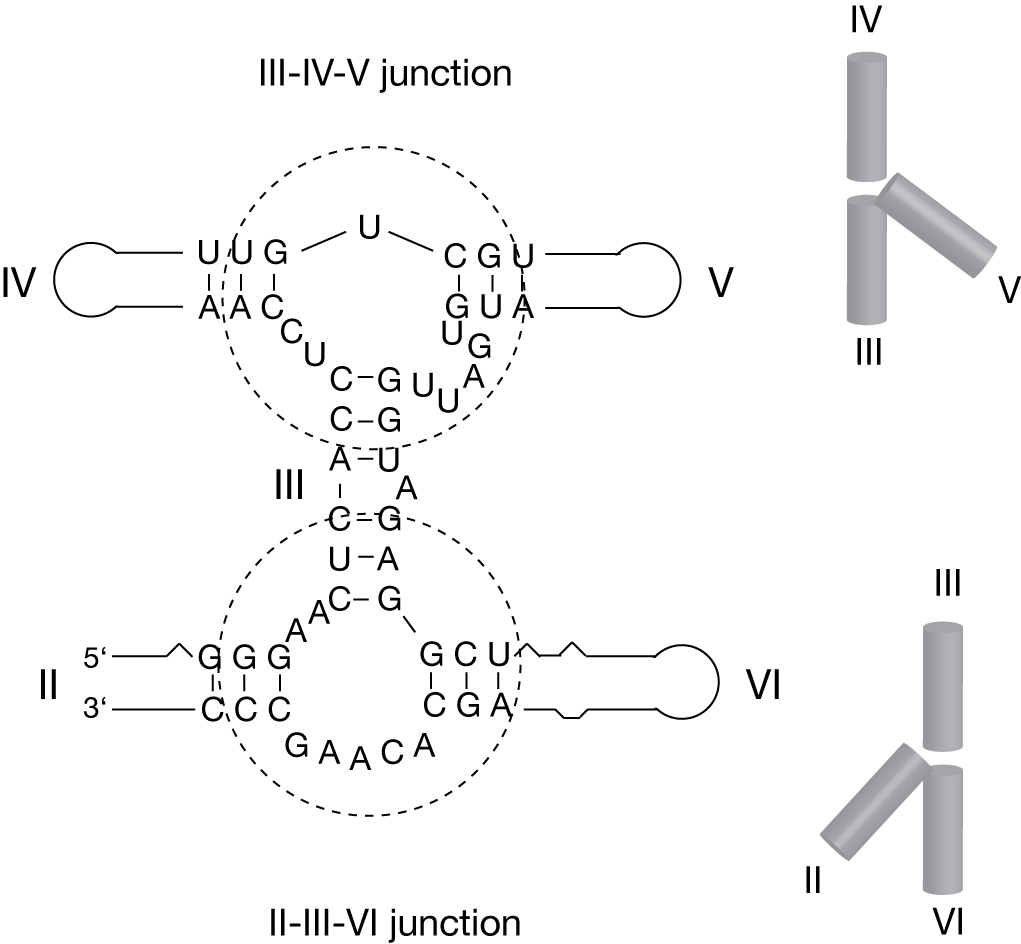
The core of the VS ribozyme is organized by two three way junctions (circled). The global shape of the junctions in the presence of metal ions is indicated.
Another elaborated three-way junction occurs in the hammerhead ribozyme. The catalytic core of the ribozyme is a HS1HS7HS3 junction, with three helices named I, II and III. Using comparative gel electrophoresis (Bassi et al., 1995) we found that in the presence of divalent metal ions, the hammerhead junction folded by the coaxial stacking of helices III on II, with an acute angle subtended between helices I and II, and this was in complete agreement with the structures observed in the crystal (Pley et al., 1994; Scott et al., 1995). However, the folding of the hammerhead junction was rather more complex than had been found for simpler junctions, such as those of the VS ribozyme. Analysis of the electrophoretic mobility patterns as a function of Mg2+ ion concentration indicated that it folded in two steps, with a distinct species existing at intermediate Mg2+ concentration in which helices II and III were coaxially stacked, but helix I was directed into the same quadrant as helix III (Bassi et al., 1995). This scheme was confirmed by steady-state FRET studies (Bassi et al., 1997), and could be rationalised in terms of the crystal structures (Pley et al., 1994; Scott et al., 1995), which showed the formation of two distinct domains within the structure. The global structure of the ribozyme fully-folded in the presence of 10 mM Mg2+ ions was in excellent agreement with that found in the crystal structures of the ribozyme (Pley et al., 1994; Scott et al., 1995). However, it was subsequently shown that tertiary contacts between elements within helices I and II that were missing in all the original constructs are important in the function of the ribozyme in physiological conditions (Canny et al., 2004; Khvorova et al., 2003), and induce a different structure in the core (Martick and Scott, 2006). We found that inclusion of these elements led to folding in a single step at significantly lower magnesium ion concentration (Penedo et al., 2004).
RNA catalysis and ribozyme mechanism
Ribozymes are RNA molecules that catalyse reactions on themselves or other molecules - enzymes that happen to be made of RNA not protein (Lilley and Eckstein, 2008). Ribozymes may have played a key role in the formation of life on the planet, whereby RNA enzymes bypassed the requirement for a translation apparatus until proteins arrived on the scene. The list of natural ribozymes is relatively short, but it includes the ribosome, RNase P and possibly the spliceosome, relics from an RNA world performing essential reactions in every eukaryotic cell.
The existence of ribozymes has been known for over a quarter of a century now, and significant progress has been made in understanding how they accomplish their function. They can achieve impressive rate accelerations (eg ~ 1011-fold for group I introns) despite having a fraction of the chemical space of proteins. RNA is limited to four rather similar heterocyclic nucleobases, and a hydroxyl group, all with rather unhelpful pKa values. There are complete or partial crystal structures for most of the known ribozymes. But while structures provide valuable insight, they seldom give us the complete story of how catalysis occurs. This requires analysis of the transition state of the reaction, not accessible by purely structural methods.
Evolution has discovered a variety of ways in which to exploit the rather limited chemical resources of RNA to achieve some impressive feats of catalysis. For example, the nucleolytic ribozyme glmS recruits a small molecule glucosamine-6-phosphate to act in general acid-base catalysis (Klein et al., 2007). This sets a fascinating precedent, with profound evolutionary implications for the origins of life; it could well be imagined that the recruitment of amino acids or peptides by ribozymes in the an RNA world might have been the first step towards the present protein-based life (Wilson and Lilley, 2009). The spliceosome has been thought likely to be a ribozyme, since its chemistry of pre-mRNA splicing is very similar to that catalysed by the group II intron, and certain domains of spliceosomal RNA show sequence similarity with the catalytic domains of the intron. However the structure of an RNase H-like domain forming part of the spliceosomal protein Prp8 (Pena et al., 2008; Ritchie et al., 2008), along with extensive genetic data, suggests that this domain serves to position the RNA within the active site. Moreover, a plausible model places the splice site adjacent to the vestigial RNase H active site. But this putative active site lacks two of the four cation binding residues observed in RNase H (Nowotny et al., 2005), and no metal ion binding has been detected (Pena et al., 2008). It therefore seems unlikely that Prp8 carries out the chemistry unaided. This raises the intriguing possibility that the spliceosome is a hybrid, with both protein and RNA domains contributing to catalysis.
We have begun to understand the mechanisms of some ribozymes but there is still much we don't know. For example, why is a particular mechanism used by a given class of ribozyme? Are multiple catalytic pathways possible?
The hairpin ribozyme
The secondary structure of the hairpin ribozyme shows it to be centred on a four-way helical junction, with arms sequentially labelled A, B, C and D (Hampel and Tritz, 1989). Adjacent arms A and B contain loops that include all but one of the nucleotides shown to be essential for catalytic activity, and the site of cleavage/ligation is located in arm A.
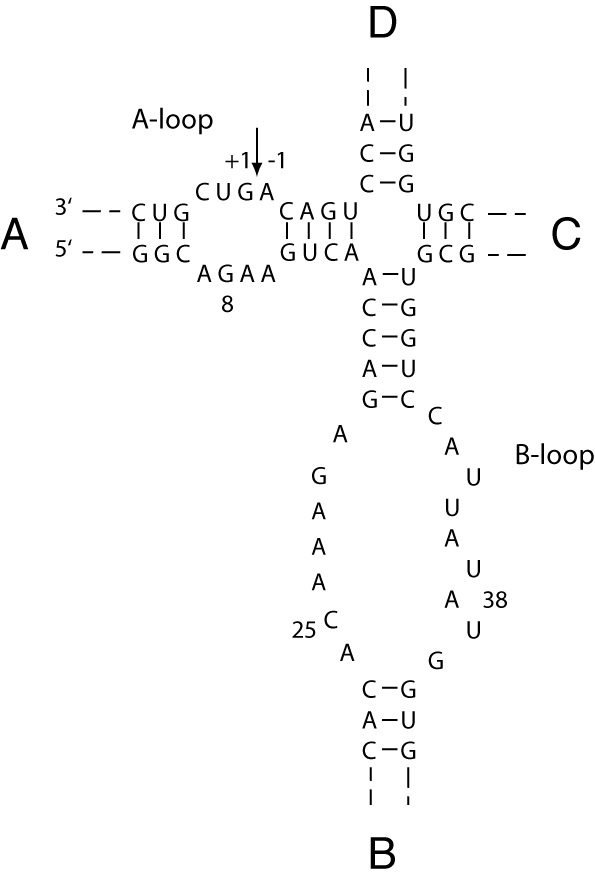
The sequence of the hairpin ribozyme in its junction form. Cleavage and ligation reactions occur at the arrowed linkage.
FRET studies with this species showed that the junction folds by coaxial stacking of helices A on D and B on C in the presence of metal ions (Murchie et al., 1998); this was the first physical evidence for loop-loop interaction. A similar analysis was performed using the hinged form lacking the C and D helices (Walter et al., 1998c). Single-molecule FRET studies of both the hinged (Zhuang et al., 2002) and four-way junction (Tan et al., 2003) forms of the ribozyme exhibit metal ion-dependent dynamics, with repeated docking and undocking of the loops. The folding of the four-way junction form occurs in two stages, with a fast (~ 100 s-1) rotation of the junction, and a slower (~3 s-1) docking process (Tan et al., 2003). The rate of docking in this form is about 500 times faster than that measured in the hinged form (Zhuang et al., 2002); folding is not rate limiting in this form of the ribozyme. While not required for catalytic activity, the junction acts as an auxiliary element that assists the folding of the ribozyme under physiological conditions (Walter et al., 1998a; Walter et al., 1998b; Walter et al., 1999), akin to the loops of the hammerhead ribozyme (Khvorova et al., 2003; Penedo et al., 2004). Crystal structures have been solved for both the junction (Rupert and Ferré-D'Amaré, 2001) and hinged form (Alam et al., 2005) of the hairpin ribozyme. These confirm the key feature of the loop-loop interaction, and the general folding principle of the junction form.
Cycles of cleavage and ligation reactions have been observed in individual hairpin ribozymes using single-molecule FRET (Nahas et al., 2004) (collaboration with Taekjip Ha). We have observed ribozyme molecules switching between two distinct dynamic states. In one state (the ligated species) the molecule remains stably docked for a period before changing into another form (the cleaved form) that undergoes rapid docking and undocking.
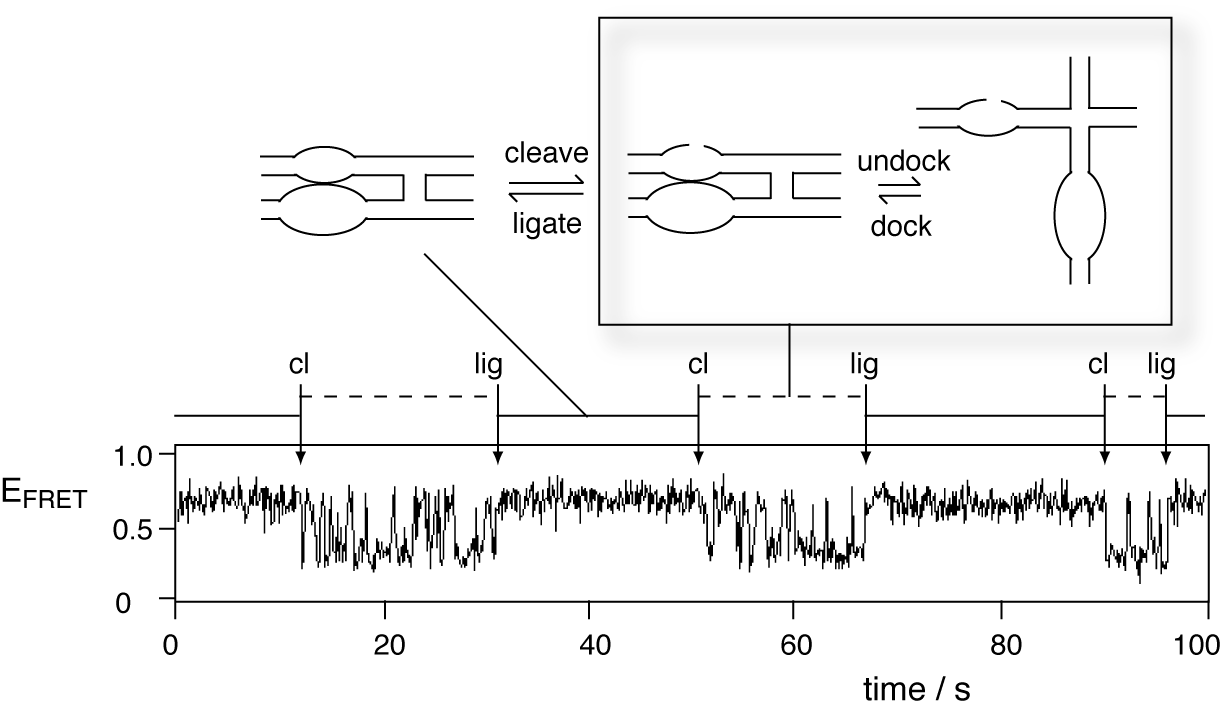
Cycles of cleavage and ligation reactions in single hairpin ribozyme molecules. The ligated state maintains a high-FRET state, while the cleaved state rapidly oscillates between high and low-FRET states.
The docking and undocking rates within the cleaved state are kCdock = 2.5 (± 0.1) s-1 and kCundock = 2.3 (± 0.1) s-1; these rates are significantly slower than the junction dynamics under the same conditions (Tan et al., 2003). The cleavage rate (kC) is ~ 0.6 min-1 in the presence of 1 mM Mg2+, while the ligation rate (kL : the rate of ligation in the docked ribozyme) is ~ 0.3 s-1. Thus the reaction is biased towards ligation, with an internal equilibrium constant of
The hinged form of the ribozyme is similarly biased towards ligation, but overall the cleavage rate is limited by docking and undocking rates (Liu et al., 2007). The bias to ligation maintains the integrity of the circular (-) strand, allowing it to act as a template for (+) strand synthesis. However, in the junction form cleavage of the concatenated product of replication is facilitated by the rapid undocking that follows cleavage. Thus the four-way junction confers the properties required for the biological function of the ribozyme.
Some time ago it was thought that bound metal ions would play the major role in accelerating the transesterification reactions of the nucleolytic ribozymes, but it is clear that this is not true for the hairpin ribozyme. Significant levels of activity could be attained in the hairpin and VS ribozymes in the presence of high concentrations of monovalent ions such as Li+ and even NH4+ (Murray et al., 1998), excluding a requirement for site-specific ion binding. Furthermore, the hairpin ribozyme remained active when Mg2+ ions were replaced by substitutionally-inert hexammine Co (III) ions (Hampel and Cowan, 1997; Nesbitt et al., 1997; Young et al., 1997), excluding a role for metal ion-bound water molecules in general acid-base catalysis. We found that the cleavage rates measured in single-molecule experiments were also found to depend very weakly on Mg2+ ion concentration (Nahas et al., 2004). This leaves nucleobase participation to provide the remaining rate acceleration. Early evidence pointed to an important role for G8 in loop A, located on the opposite strand from the scissile phosphate. Substitution by other nucleotides led to rates of cleavage being reduced by several orders of magnitude in both the natural (Wilson et al., 2001) and hinged form (Pinard et al., 2001) of the ribozyme, without affecting ion-induced folding. Thomas and Perrin (Thomas and Perrin, 2006) demonstrated alkylation of G8 when the 2'-OH nucleophile was substituted by bromoacetamide, further supporting the direct participation of this nucleobase.
Ferré d'amaré and colleagues (Rupert et al., 2002) provided into the nature of the active site by a crystal structure of a modified hairpin ribozyme in which the scissile phosphate replaced by vanadate to provide a model of a pentacoordinate phosphorane transition state. It was found that G8 is hydrogen bonded to the 2'-O and the proS non-bridging O of the scissile phosphate, well positioned to participate in the catalytic chemistry. The structure also revealed the presence of a second nucleotide juxtaposed with the scissile phosphate - the nucleobase of A38 (contributed by loop B) formed hydrogen bonds to the 5'-O and the proR O. Removal of the nucleobase at position 38 led to a 10,000-fold loss of activity (Kuzmin et al., 2005), while ligation activity was shown to be sensitive to substitution by inclusion of modified nucleotides in nucleotide analog interference mapping experiments (Ryder et al., 2001). The nucleobases of G8 and A38 should provide some stabilization of the transition state, and consequently a potential rate enhancement. However, they are also well placed to participate in coordinated general acid-base catalysis. G8 is positioned to deprotonate the 2'-O, while A38 could protonate the 5'-oxyanion leaving group in the cleavage reaction. By the principle of microscopic reversibility, G8 and A38 would act as general acid and base respectively in the ligation reaction. The pH dependence of hairpin ribozyme rates are consistent with the participation of two bases with pKa values of ~ 6.2 and > 9.5 (Bevilacqua, 2003; Kuzmin et al., 2004; Nahas et al., 2004; Pinard et al., 2001). The lower value is within the range of an adenine base with a pKa perturbed by an electronegative environment.
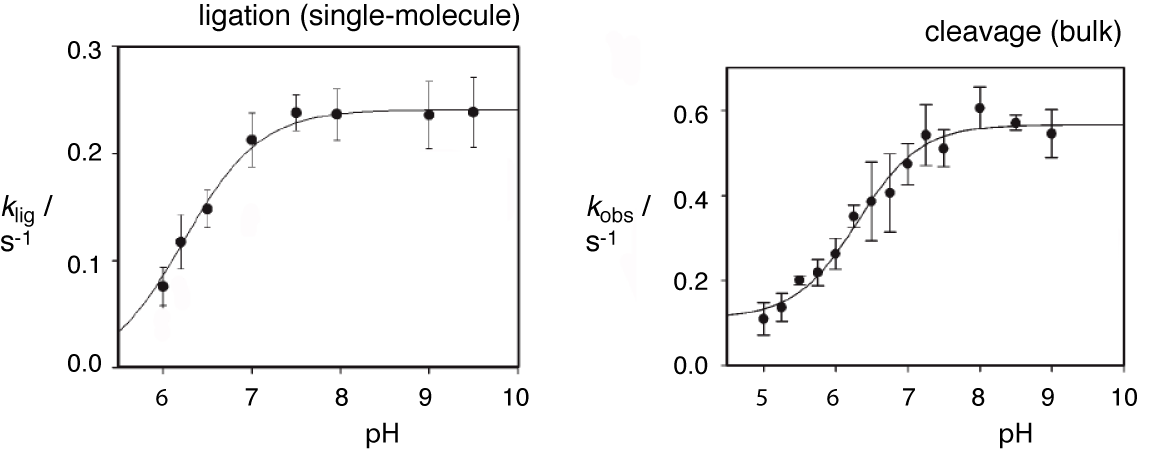
The pH dependence of ligation (left, measured from single-molecule experiments like those above) and cleavage (right, measured in conventional bulk experiments) reactions in the hairpin ribozyme. Under these conditions a single ionization is observed, corresponding to pKa values of 6.2 ± 0.1 and 6.3 ± 0.1 respectively.
With Shinya Harusawa (Osaka) we have introduced a new chemo-genetic strategy to investigate general acid-base catalysis in ribozymes in which imidazole base is introduced as a pseudo-nucleoside by chemical synthesis (Araki et al., 2004). We have now created a series of imidazole nucleosides (as phosphoramidites) with zero, two and three carbon atoms linking the imidazole heterocycle to the ribose.
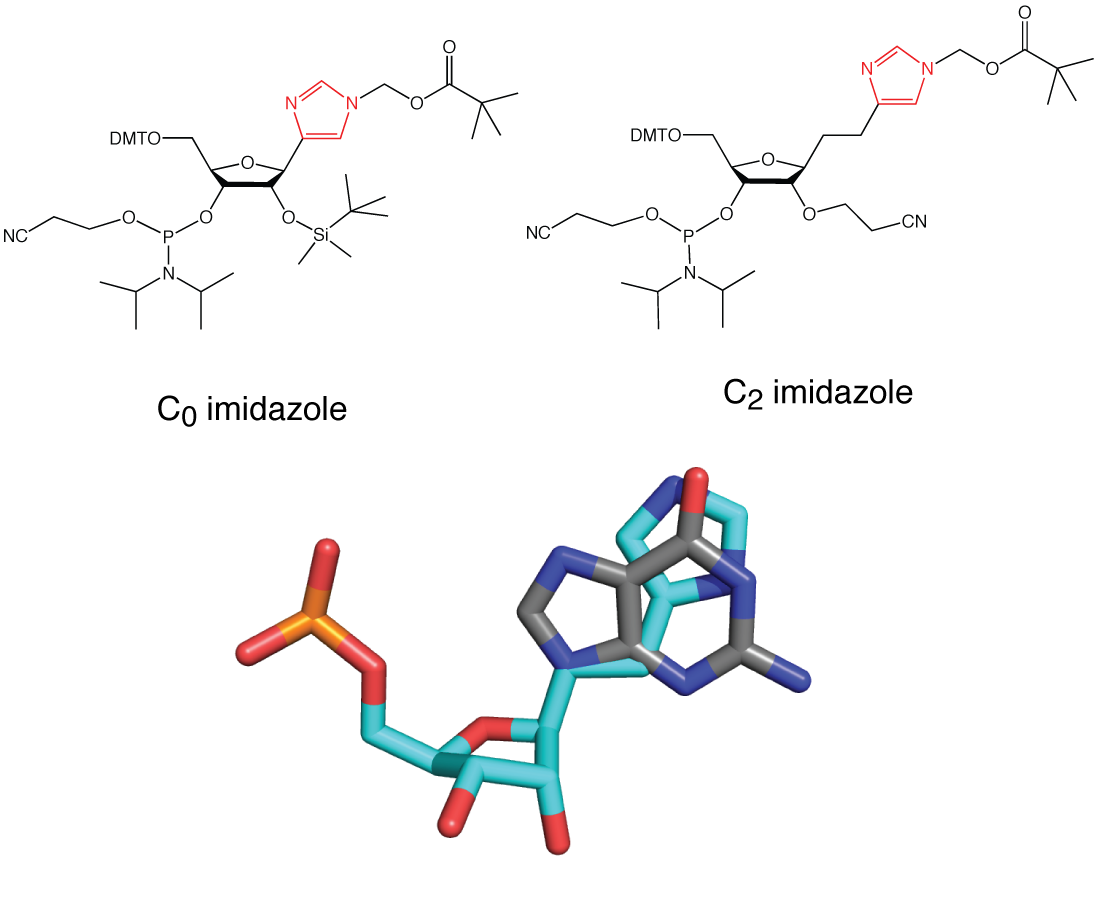
A series of imidazole nucleosides. Top shows the C0 (left) and C2 (right) compounds as phosphoramidites suitable for incorporation into RNA by chemical synthesis. At the bottom is show a superposition of the C2-linked imidazole with guanine.
Using this approach, we generated a hairpin ribozyme in which a C0-linked imidazole nucleoside was introduced at position 8 in place of guanine (Wilson et al., 2006). The substitution did not perturb folding, and the modified ribozyme was active in both cleavage and ligation. The cleavage product was indistinguishable from that of the natural ribozyme. The rate of cleavage measured at pH 7 in the presence of 10 mM Mg2+ ions at 25°C was ten-fold faster than that for the G8U variant (Wilson et al., 2001). But importantly, the modified ribozyme gave a bell-shaped pH profile. These data were fitted to a model based on two proton transfers, giving estimated pKa values of 5.7 ± 0.1 and 7.7 ± 0.1. The higher pKa should be that of the imidazole. A bell-shaped pH dependence was also observed for the ligation reaction, with pKa values of 6.1 ± 0.3 and 6.9 ± 0.3.
The Varkud Satellite ribozyme
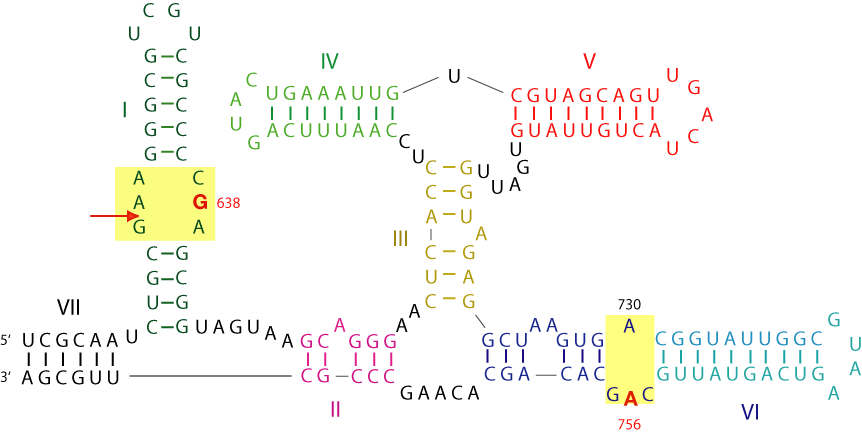
The sequence of the VS ribozyme (Beattie et al., 1995) in its most complete form including helix VII. Cleavage or ligation occurs at the arrowed position. Helix I can be cleaved as a separate stem-loop structure, catalyzed by a trans-acting ribozyme comprising helices II - VI. The two areas critical to the activity of the ribozyme are shaded yellow, and the key nucleotides A756 and G638 are highlighted in red.
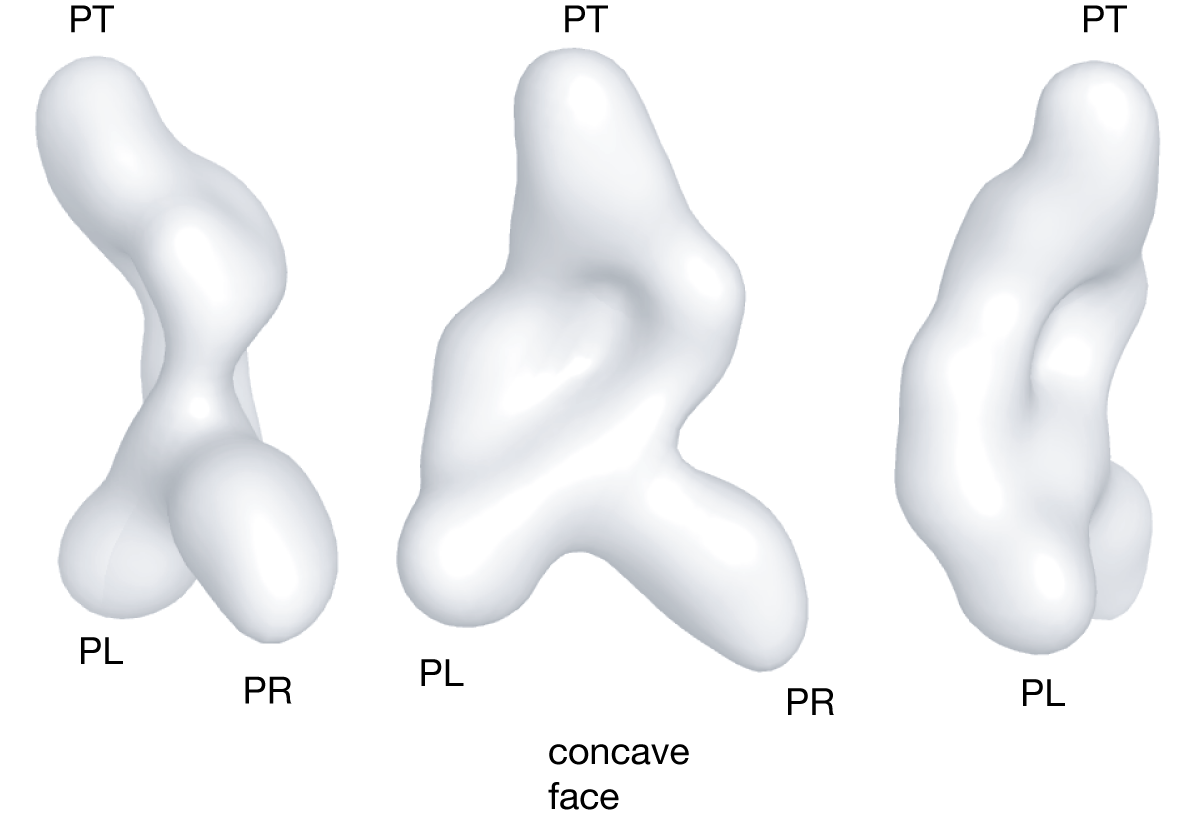
Initial biophysical studies focussed on the II-III-VI and III-IV-V junctions, as explained above using a combination of comparative gel electrophoresis, FRET and homology-based modeling (Lafontaine et al., 2001a; Lafontaine et al., 2002). From this we proposed a core structure based upon the coaxial alignment of helices IV, III and VI, with helices II and V directed laterally. The dihedral angle between helices II and V was determined by an electrophoretic technique. We suggested that the probable location of helix I was in the cleft formed between helices II and VI, so that it could be connected to helix II and make the loop-loop interaction with helix V (Lafontaine et al., 2002). We used small-angle X-ray scattering (SAXS) in solution to study the component junctions, the trans-acting form (helices II-VI) and the complete ribozyme comprising helices I through VII (Lipfert et al., 2008). Radii of gyration and maximum chord length measurements confirmed the probable coaxial alignment of helices IV-III-VI. Reconstructions based on the full scattering curves generated a family of closely-related structures that were broadly in agreement with the earlier model. The overall structure is rather flat, with helix-width protrusions at three corners, and a helical ridge running across the face. These features enabled us to assign the features of the low-resolution electron density envelope to the known helical sections of the ribozyme. Further details on this process can be found at VS Mini Site.
| |
The structure of the VS ribozyme as determined by SAXS studies. On the left is shown three views of the electron density map for the complete VS ribozyme RNA. A model of the ribozyme fitting to the density map is shown on the right hand side. The orientation fits to the central view of the density.
The key catalytic components of the VS ribozyme have been identified by nucleotide substitution. Sequence changes in the internal loop of helix VI (the A730 loop) have no effect on folding, yet single-base changes introduced into this loop result in loss of cleavage activity in trans (Lafontaine et al., 2001b) and in cis (Sood and Collins, 2002). The A730 loop had been found to be sensitive to ethylation interference, and contained sites of interference by phosphorothioate incorporation, and suppression by thiophilic metal ions (Sood et al., 1998). Sequence changes in the loop also affected ligation activity (McLeod and Lilley, 2004), which was also very sensitive to the introduction of nucleotide analogs (Jones and Strobel, 2003). The scissile phosphate is readily juxtaposed with the A730 loop in our SAXS-derived model of the ribozyme (Lipfert et al., 2008). Within the A730 loop, substitution of A756 by G, C or U leads to a large loss of cleavage (Lafontaine et al., 2001b) and ligation activity (McLeod and Lilley, 2004). These changes have only a small effect on substrate binding affinity (≤ 5 fold weaker Kd), and most of the effect arises from a reduced rate of central conversion of the substrate into product (Lafontaine et al., 2001b). A variant ribozyme with an imidazole nucleotide covalently located at position 756 gave significant rates of cleavage and ligation reactions (Zhao et al., 2005).
We identified a second nucleobase involved in catalysis (Wilson et al., 2007). Replacement of G638 in the internal loop of the substrate by any other nucleotide resulted in an impairment of cleavage rate by four orders of magnitude, while substrate folding was not perturbed. The variant substrates bind the ribozyme with similar affinity, acting as competitive inhibitors of the natural sequence substrate. Thus a G638A substrate binds to the ribozyme in an apparently normal manner, but is catalytically impaired.
These studies collectively point to an involvement of two key nucleobases, A756 and G638 in the catalytic mechanism of the VS ribozyme. An intimate association of the A730 loop and the internal loop of substrate helix I would bring these nucleotides and the scissile phosphate into juxtaposition, as indicated by the SAXS-derived model of the ribozyme (Lipfert et al., 2008).
The catalytic mechanism of the VS ribozyme
A756 and G638 are very important to the catalytic mechanism of the VS ribozyme. They could have a number of possible roles. They might stabilize the transition state by hydrogen bonding the phosphorane, or a positively-charged protonated adenine base might provide electrostatic stabilization of the dianionic transition state. But present evidence points towards general acid-base catalysis by these nucleobases.
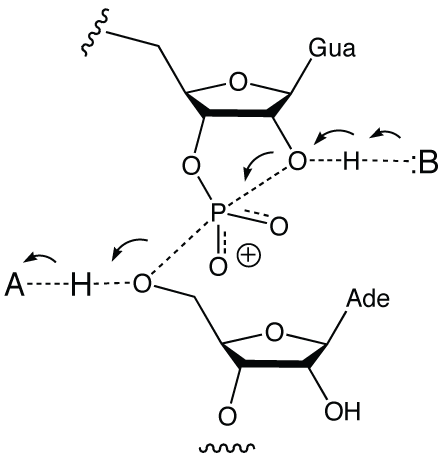
A general mechanism of ribozyme cleavage of RNA using general acid base catalysis.
Kinetic isotope effects in a fast, cis-acting form of the VS ribozyme show that proton transfer occurs in the transition state of the cleavage reaction (Smith and Collins, 2007). Both A756 and G638 can be replaced with imidazole nucleosides with retention of some activity (Araki et al., 2009; Zhao et al., 2005). This catalytic strategy requires that the acid be protonated and the base unprotonated at the outset of the reaction. The observed rate of reaction will be given by
where fA and fB are the fractions of protonated acid and unprotonated base respectively, and kcat is the rate of cleavage catalyzed by the ribozyme in the correct state of protonation. fA and fB can be calculated for any pH if pKa values are assumed, and thus the pH dependence of cleavage rate simulated (Bevilacqua, 2003). We measured the pH dependence of the cleavage reaction in the presence of a high concentration of Mg2+ ions, and obtained a bell-shaped pH dependence (Wilson et al., 2007).
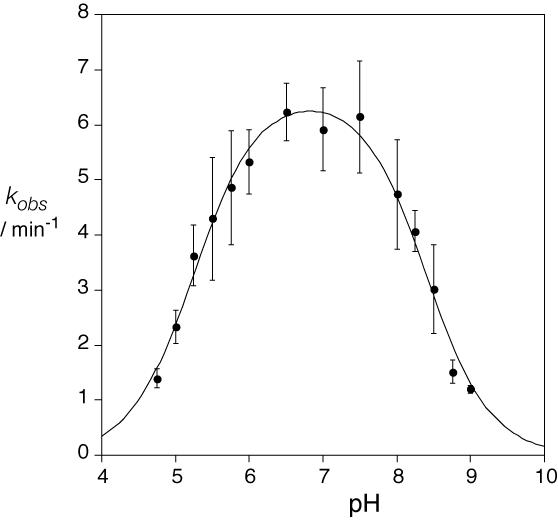
The pH dependence of the rate of cleavage by the VS ribozyme. The data were fitted to a double-ionization model, with apparent pKa values of 5.2 and 8.4.
The lower value is very much in line with an adenine in an electronegative environment, while the upper value would be consistent with a guanine base with a reduced pKa (perhaps by proximity to metal ions). Smith and Collins (Smith and Collins, 2007) also obtained a bell-shaped pH dependence for cleavage using the fast-cleaving cis-acting form of the ribozyme, with pKa values of 5.8 and 8.3. Substitution by 2,6-diaminopurine (with a normal pKa of 5.1) at position 638 gave a rate of cleavage that was reduced 1000-fold relative to the natural sequence at pH 8, but significantly restored at lower pH, corresponding to a pKa of 5.6. The rate was log-linear with pH over the range 6 - 8, with a gradient of one. This indicates that general acid-base catalysis contributes over two orders of magnitude to the catalytic power of the VS ribozyme.
Present evidence indicates that the base in the cleavage reaction is G638, and the acid is A756. In collaboration with Dr Joe Piccirilli (University of Chicago) we introduced a 5'-phosphorothiolate (5'-PS) substitution at the scissile phosphate (Wilson et al., 2010). This restored the activity of an A756G substituted-ribozyme, consistent with the requirement for A756 acting as a general acid in the oxy form. The pH dependence of the 5'-PS ribozyme acting in the cleavage reaction was no longer bell-shaped, but reached a plateau at around neutrality, as expected if the reaction requires only general base catalysis by the nucleobase at position 638.

The proposed mechanism for the VS ribozyme cleavage reaction.
In this mechanism the proton is removed from the 2'-O nucleophile by G638, and the 5'-oxyanion leaving group is protonated by A756. By the principle of microscopic reversibility, in the reverse (ligation) reaction G638 will act as an acid (protonating the 2'-oxyanion leaving group) and A756 will be the base (removing the proton from the 5'-O nucleophile).
Mechanistic similarities between the hairpin and VS ribozymes
We have argued that the catalytic mechanisms of the hairpin and VS ribozymes are likely to be closely similar (Wilson and Lilley, 2011). Both appear to work by general acid-base catalysis, based on a guanine acting as the base in cleavage (G8 and 638 in the hairpin and VS respectively), and an adenine as the general acid (A38 and 756 respectively). Further experiments are in hand to test this proposal.
The kink-turn in RNA
The kink-turn (usually abbreviated to k-turn) is a widespread structural motif in RNA, first noted as a repeated structural element in the large ribosomal subunit by the Steitz lab (Klein et al., 2001). It introduces a very tight kink into the axis of helical RNA. It clearly plays an important structural role in RNA, and is significant in many aspects of RNA function including translation, modification and splicing, as well as genetic regulation. Many k-turns are bound by proteins in their natural state, at least some of which stabilize the kinked geometry.
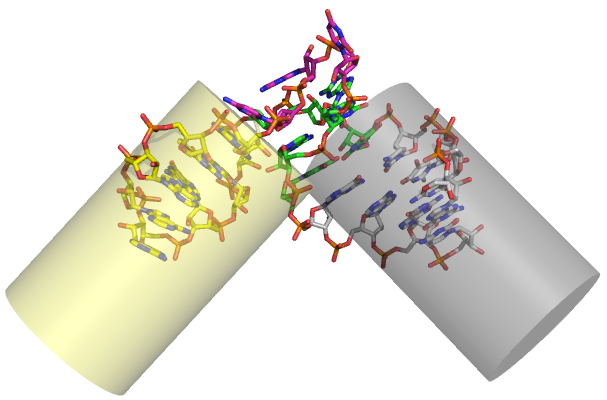
The canonical k-turn comprises a three-nucleotide bulge flanked on its 3' side by A•G and G•A basepairs (the N-C stem), and on its 5' side by a section of regular basepairing (the C stem) (Klein et al., 2001). Kt-7 of the 23S rRNA of the H. marismortui ribosome is close to the consensus sequence for k-turns. The structure introduces a pronounced kink in the RNA (hence its name), with an included angle between the axes of ~ 60°. The minor grooves of the two helices are juxtaposed, and the conformation is stabilized by interactions between the stacked adenosines of the A•G mismatches and the C stem, and by stacking of the 5' and central bases of the bulge on the ends of the C and N-C stems respectively.
| |
The k-turn. motif. Left : the sequence of Kt-7, which is close to the archetypal k-turn. Right : the structure of a k-turn taken from the 50S ribosomal subunit.
We have created an internet site dedicated to the structure and folding of k-turns. This is an on-line data base of sequence and structural information (Schroeder et al., 2010). Three-dimensional views of all available k-turns can be viewed, manipulated and compared.
We have shown that k-turns are not tightly kinked in the absence of metal ions, but adopt the kinked geometry on addition of Mg2+ ions (Goody et al., 2004), or on binding proteins of the L7Ae family (Turner et al., 2005). L7Ae binds with very picomolar affinity (Turner and Lilley, 2008), and is a good example of induced fit. We have defined the key hydrogen bonding interactions that are important in stabilizing the kinked conformation (Liu and Lilley, 2007; Schroeder and Lilley, 2009; Turner and Lilley, 2008). Additionally, tertiary interactions can also stabilize kink-turn structure, as exemplified by the SAM-I riboswitch (Schroeder et al., 2011).
Development of methods for the study of branched nucleic acids
In general, branched nucleic acids are too large for NMR, and do not readily crystallize. We have consequently been forced to improvise, and exploit other biophysical methods where possible. The simplest of these is perhaps comparative gel electrophoresis (Lilley, 2009a), and was developed from experiments in this laboratory (Duckett et al., 1988; Gough and Lilley, 1985) and that of Paul Hagerman (Cooper and Hagerman, 1987). Despite the lack of a solid theoretical basis, comparative gel electrophoresis is relatively easy to perform and extremely powerful. It is frequently where we begin any new study, and it can sometimes resolve questions of ambiguity. It successfully solved the structure of the four-way junction (Duckett et al., 1988) more than a decade before the crystallographers (who were not setting out to solve that problem anyway !), and indeed to date it has never mislead (Lilley, 2008). More recently, we have found that SAXS can be very powerful for the analysis of folded RNA structures (Lipfert et al., 2008). But a consistent thread running though the studies of nucleic acid conformation in this laboratory for many years has been the application of FRET in various forms.
Fluorescence resonance energy transfer
The demonstration of the structure of the four-way DNA junction using FRET between different pairs of helices terminally labelled with fluorescein and rhodamine (Murchie et al., 1989) established a methodology that has been exploited many times since (Lilley, 2009b). These studies have included both steady-state and time-resolved measurements in bulk, and more recently the extension to single molecules.
The cyanine fluorophores have become very popular in single-molecule studies of nucleic acids, and we have used them extensively for this purpose. We have shown that both Cy3 (Norman et al., 2000) and Cy5 (Iqbal et al., 2008b) stack upon the ends of double-helical DNA very much in the manner of an additional basepair when they are attached to 5'-termini via C3 linkers, as they are when coupled as phosphoramidites during synthesis. These structures are available at Cy3-Cy5 Mini Site.
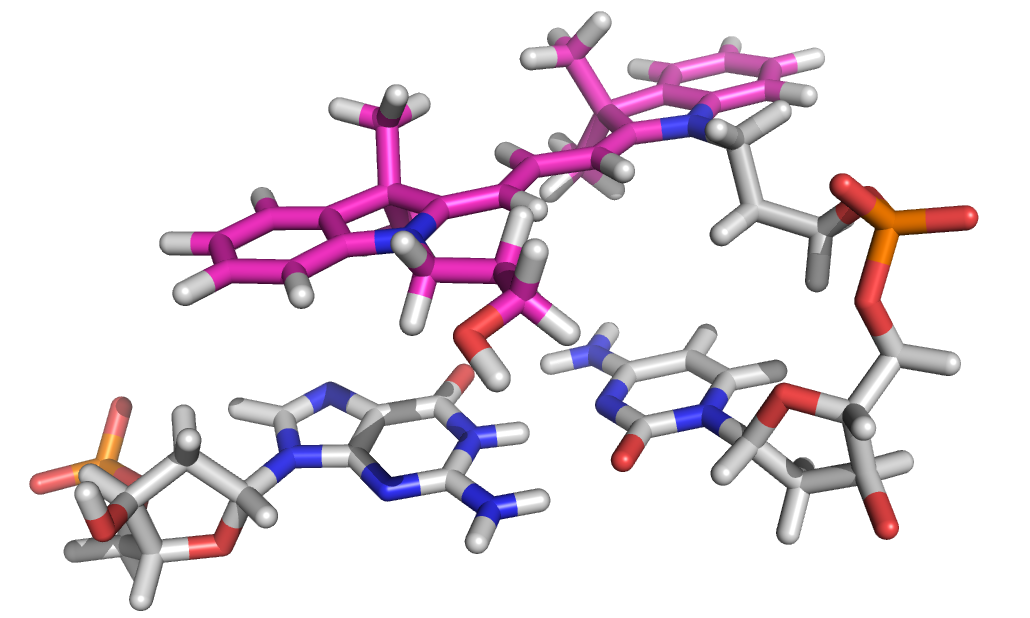
NMR structure of Cy3 (magenta) attached to the end of double-stranded DNA (Norman et al., 2000). A rather similar structure of Cy5 bound to DNA has recently been determined (Iqbal et al., 2008b).
The relative immobilization of the fluorophores could result in a significant orientation dependence of dipolar energy transfer between them. FRET arises from the dipolar coupling between the oscillating transition dipoles of the donor and acceptor fluorophores. The efficiency of energy transfer (EFRET) is given by
where rDA is the separation between the donor and acceptor. ki are the rates of all possible mechanisms for the relaxation of the excited state. R0 is the characteristic Förster length for a given donor-acceptor pair of fluorophores, given by :
where the units of R0 and the wavelength λ are in cm. ΦD is the fluorescent quantum yield of the donor in the absence of the acceptor and N is the Avogadro number. n is the refractive index of the medium in which the electric fields of the transition dipoles extend; a value of 1.33 is appropriate for the aqueous medium of biological macromolecules. J( λ ) is the normalized spectral overlap between donor emission and acceptor absorption, given by :
where φD is the spectral shape function for the donor emission and εA is that for acceptor absorption (in M-1 cm-1). κ is a parameter that depends on the relative orientation of the donor and acceptor transition moments, given by :
where and
are the donor and acceptor transition dipole moment vectors,
is the vector between their centers, and the angles ΘT, ΘD and ΘA are defined below.
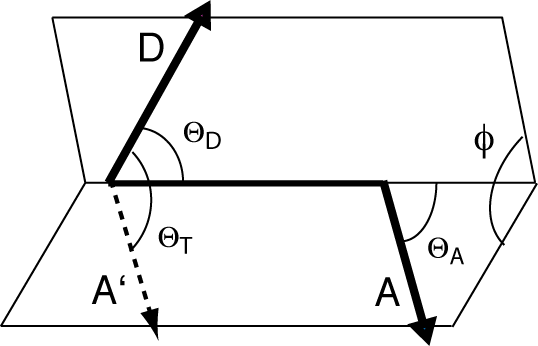
Parameters describing the relative orientation between donor (D) and acceptor (A) transition moment dipole vectors.
κ2 can take values between 0 and 4. If the value is not known, it becomes difficult to extract distances from FRET efficiencies. We made a demonstration of this using a series of DNA and RNA.DNA duplexes of lengths between 10 and 24 bp, with Cy3 and Cy5 attached at the 5' termini (Iqbal et al., 2008a). The fluorophores (and hence their transition dipoles) will sit with their planes approximately parallel and coaxial, but the angle between their transitions moments (that are approximately directed along the polymethyne linkers between the two indole rings (Iqbal et al., 2008b)) will depend upon the length of the helix and its periodicity. κ2 should be maximal ( κ2 ~ 1) when the transition dipoles are parallel, and minimal ( κ2 ~ 0) when they are perpendicular. Thus it would be expected that EFRET should be modulated with twice the periodicity of the helix.

FRET efficiency as a function of helix length for DNA (squares) and RNA.DNA (circles) duplexes (Iqbal et al., 2008a). The data were simulated (lines) on the basis of standard B- and A-form geometries respectively, with the fluorophores allowed to undergo a Gaussian distribution of lateral mobility with respect to each other.
For both DNA and RNA.DNA duplexes the FRET efficiency falls with length, but with a pronounced modulation. EFRET is modulated with twice the helical periodicity; for example for RNA there are clear maxima at 11 and 17 bp, with minima at 15 and 21 bp. The DNA helix exhibits the same behavior, but with a phase shift indicative of a more tightly-wound B-form helix. This is just the behavior expected for orientation dependence of FRET. Yet the modulation is clearly incomplete (EFRET does not become zero at the minima), suggesting that the orientation is subject to averaging by a combination of lateral mobility of the fluorophores on the end of the helix, and torsional flexibility of the helices. The data were well-simulated by a model based on the geometric properties of standard B- and A-form helices for DNA and RNA.DNA, and the positions of the fluorophores determined by NMR with a significant lateral averaging (Iqbal et al., 2008a), with or without a fraction of unstacked fluorophore as indicated by time-resolved studies (Iqbal et al., 2008a; Sanborn et al., 2007). This is a nice confirmation of the expected orientational dependence of Förster resonance energy transfer, but also a warning that such effects could significantly affect the interpretation of FRET data in terms of distance with a commonly-used pair of fluorophores. Yet it also means that we should be able to extract useful angular information from FRET using Cy3 and Cy5 in nucleic acids. And if we understand the orientational dependence better, then we can also obtain more reliable distance information.
We have recently generated a proof-of-principle that we can obtain orientational information by FRET. We repeated the analysis of a DNA duplex series using Cy3 and Cy5 sulfoindocarbocyanine fluorophores linked via 13-atom tethers to the 5'-O of the DNA, in contrast to the C3 linkers used above. We found that EFRET was modulated with distance as before, despite the longer tether, so that the terminal stacking of the cyanine fluorophores is clearly intrinsic, not a result of the linkage. However, interestingly a significant phase shift was observed relative to the data with the shorter tether (Ouellet et al., 2011), which indicated that each fluorophore is rotated by 30° (relative to the short-tether data), so that the long axis of each fluorophore is approximately parallel to that of the terminal basepair. This conclusion emerging from the FRET data was subsequently confirmed exactly by NMR study (Urnavicius et al., 2012). Thus the FRET really did get it right - it can provide reliable information on orientation.
References
Alam, S., Grum-Tokars, V.,
Krucinska, J., Kundracik, M.L. and Wedekind, J.E. (2005) Conformational
heterogeneity at position U37 of an all-RNA hairpin ribozyme with implications
for metal binding and the catalytic structure of the S-turn. Biochemistry, 44, 14396-14408.
Araki, L., Harusawa, S., Yamaguchi, M.,
Yonezawa, S., Taniguchi, N., Lilley, D.M.J., Zhao, Z. and Kurihara, T. (2004)
Synthesis of C4-linked imidazole ribonucleoside phosphoramidite with
pivaloyloxymethyl (POM) group. Tetrahedron
Lett., 45, 2657-2661.
Araki, L., Morita, K., Yamaguchi, M., Zhao, Z.,
Wilson, T.J., Lilley, D.M.J. and Harusawa, S. (2009) Synthesis of novel
C4-linked C2-imidazole ribonucleoside phosphoramidite, and its
application to probing the catalytic mechanism of a ribozyme. . J. Org. Chem., 74, 2350-2356.
Ariyoshi, M., Vassylyev, D.G., Iwasaki, H.,
Nakamura, H., Shinagawa, H. and Morikawa, K. (1994) Atomic structure of the
RuvC resolvase: A Holliday junction-specific endonuclease from E.coli. Cell, 78, 1063-1072.
Bailly, A.P., Freeman, A., Hall, J., Déclais,
A.-C., Alpi, A., Lilley, D.M.J., Ahmed, S. and Gartner, A. (2010) The Gen-1
resolvase links DNA damage signaling to DNA double-strand break repair. PLoS Genetics, 6, e1001025.
Bassi, G., Mřllegaard, N.E., Murchie, A.I.H.,
von Kitzing, E. and Lilley, D.M.J. (1995) Ionic interactions and the global
conformations of the hammerhead ribozyme. Nature
Struct. Biol., 2, 45-55.
Bassi, G.S., Murchie, A.I.H., Walter, F.,
Clegg, R.M. and Lilley, D.M.J. (1997) Ion-induced folding of the hammerhead
ribozyme: a fluorescence resonance energy transfer study. EMBO J., 16, 7481-7489.
Beattie, T.L., Olive, J.E. and Collins, R.A.
(1995) A secondary-structure model for the self-cleaving region of Neurospora VS RNA. Proc. Natl. Acad. Sci. USA, 92,
4686-4690.
Bevilacqua, P.C. (2003) Mechanistic
considerations for general acid-base catalysis by RNA: revisiting the mechanism
of the hairpin ribozyme. Biochemistry,
42, 2259-2265.
Biertümpfel, C., Yang, W. and Suck, D. (2007)
Crystal structure of T4 endonuclease VII resolving a Holliday junction. Nature, 449, 616-620.
Bond, C.S., Kvaratskhelia, M., Richard, D.,
White, M.F. and Hunter, W.N. (2001) Structure of Hjc, a Holliday junction
resolvase, from Sulfolobus solfataricus. Proc.
Natl. Acad. Sci. USA, 98,
5509-5514.
Canny, M.D., Jucker, F.M., Kellogg, E.,
Khvorova, A., Jayasena, S.D. and Pardi, A. (2004) Fast cleavage kinetics of a
natural hammerhead ribozyme. J. Amer.
Chem. Soc., 126, 10848-10849.
Ceschini, S., Keeley, A., McAlister, M.S.B.,
Oram, M., Phelan, J., Pearl, L.H., Tsaneva, I.R. and Barrett, T.E. (2001)
Crystal structure of the fission yeast mitochondrial Holliday junction
resolvase Ydc2. EMBO J., 20, 6601-6611.
Clegg, R.M., Murchie, A.I.H., Zechel, A.,
Carlberg, C., Diekmann, S. and Lilley, D.M.J. (1992) Fluorescence resonance
energy transfer analysis of the structure of the four-way DNA junction. Biochemistry, 31, 4846-4856.
Cooper, J.P. and Hagerman, P.J. (1987) Gel
electrophoretic analysis of the geometry of a DNA four-way junction. J. Molec. Biol., 198, 711-719.
Déclais, A.-C., Hadden, J.M., Phillips, S.E.V.
and Lilley, D.M.J. (2001) The active site of the junction-resolving enzyme T7
endonuclease I. J. Molec. Biol., 307, 1145-1158.
Déclais, A.-C. and Lilley, D.M.J. (2000)
Extensive central disruption of a four-way junction on binding CCE1 resolving
enzyme. J. Molec. Biol., 296, 421-433.
Déclais, A.C. and Lilley, D.M.J. (2008) New
insight into the recognition of branched DNA structure by junction-resolving
enzymes. Curr. Opin. Struct. Biol., 18, 86-95.
Déclais, A.C., Liu, J., Freeman, A.D.J. and
Lilley, D.M.J. (2006) Structural recognition between a four-way DNA junction
and a resolving enzyme J. Molec. Biol.,
359, 1261-1276.
Duckett, D.R., Murchie, A.I.H., Diekmann, S.,
von Kitzing, E., Kemper, B. and Lilley, D.M.J. (1988) The structure of the
Holliday junction and its resolution. Cell,
55, 79-89.
Duckett, D.R., Murchie, A.I.H. and Lilley,
D.M.J. (1995) The global folding of four-way helical junctions in RNA,
including that in U1 snRNA. Cell, 83, 1027-1036.
Eichman, B.F., Vargason, J.M., Mooers, B.H.M.
and Ho, P.S. (2000) The Holliday junction in an inverted repeat DNA sequence:
Sequence effects on the structure of four-way junctions. Proc. Natl. Acad. Sci. USA, 97,
3971-3976.
Fogg, J.M., Kvaratskhelia, M., White, M.F. and
Lilley, D.M.J. (2001) Distortion of DNA junctions imposed by the binding of
resolving enzymes: A fluorescence study. J.
Molec. Biol., 313, 751-764.
Fogg, J.M. and Lilley, D.M.J. (2001) Ensuring
productive resolution by the junction-resolving enzyme RuvC: Large enhancement
of second-strand cleavage rate. Biochemistry,
39, 16125-16134.
Fogg, J.M., Schofield, M.J., Déclais, A.-C. and
Lilley, D.M.J. (2000) The yeast resolving enzyme CCE1 makes sequential
cleavages in DNA junctions within the lifetime of the complex. Biochemistry, 39, 4082-4089.
Giraud-Panis, M.-J.E. and Lilley, D.M.J. (1997)
Near-simultaneous DNA cleavage by the subunits of the junction-resolving enzyme
T4 endonuclease VII. EMBO J., 16, 2528 - 2534.
Goody, T.A., Melcher, S.E., Norman, D.G. and
Lilley, D.M.J. (2004) The kink-turn motif in RNA is dimorphic, and metal ion
dependent. RNA, 10, 254–264.
Gough, G.W. and Lilley, D.M.J. (1985) DNA
bending induced by cruciform formation. Nature,
313, 154 - 156.
Grainger, R.J., Murchie, A.I.H. and Lilley,
D.M.J. (1998) Exchange between stacking conformers in a four-way DNA junction. Biochemistry, 37, 23-32.
Hadden, J.M., Convery, M.A., Déclais, A.-C.,
Lilley, D.M.J. and Phillips, S.E.V. (2001) Crystal structure of the Holliday
junction-resolving enzyme T7 endonuclease I at 2.1 Ĺ resolution. Nature Struct. Biol., 8, 62-67.
Hadden, J.M., Déclais, A.-C., Carr, S., Lilley,
D.M.J. and Phillips, S.E.V. (2007) The structural basis of Holliday junction
resolution by T7 endonuclease I. Nature,
449, 621-624.
Hadden, J.M., Déclais, A.-C., Phillips, S.E.V.
and Lilley, D.M.J. (2002) Metal ions bound at the active site of the
junction-resolving enzyme T7 endonuclease I. EMBO J., 21, 3505-3515.
Hampel, A. and Cowan, J.A. (1997) A unique
mechanism for RNA catalysis: the role of metal cofactors in hairpin ribozyme
cleavage. Chem. & Biol., 4, 513-517.
Hampel, A. and Tritz, R. (1989) RNA catalytic
properties of the minimum (-)sTRSV sequence. Biochemistry, 28,
4929-4933.
Hohng, S., Wilson, T.J., Tan, E., Clegg, R.M.,
Lilley, D.M.J. and Ha, T. (2004) Conformational flexibility of four-way junctions
in RNA. J. Molec. Biol., 336, 69-79.
Hohng, S., Zhou, R., Nahas, M.K., Yu, J.,
Schulten, K., Lilley, D.M.J. and Ha, T. (2007) Fluorescence-force spectroscopy
maps two-dimensional reaction landscape of the Holliday junction. Science, 318, 279-283.
Holliday, R. (1964) A mechanism for gene
conversion in fungi. Genet. Res,, 5, 282-304.
Ip, S.C., Rass, U., Blanco, M.G., Flynn, H.R.,
Skehel, J.M. and West, S.C. (2008) Identification of Holliday junction
resolvases from humans and yeast. Nature,
456, 357-361.
Iqbal, A., Arslan, S., Okumus, B., Wilson,
T.J., Giraud, G., Norman, D.G., Ha, T. and Lilley, D.M.J. (2008a) Orientation
dependence in fluorescent energy transfer between Cy3 and Cy5
terminally-attached to double-stranded nucleic acids. Proc. Natl. Acad. Sci. USA, 105,
11176-11181.
Iqbal, A., Wang, L., Thompson, K.C., Lilley,
D.M.J. and Norman, D.G. (2008b) The structure of cyanine 5 terminally attached
to double-stranded DNA: implications for FRET studies. Biochemistry, 47,
7857-7862.
Jones, F.D. and Strobel, S.A. (2003) Ionization
of a critical adenosine residue in the Neurospora
Varkud Satellite ribozyme active site. Biochemistry,
42, 4265-4276.
Joo, C., McKinney, S.A., Lilley, D.M.J. and Ha,
T. (2004) Exploring rare conformational species and ionic effects in DNA
Holliday junctions using single-molecule spectroscopy. J. Molec. Biol., 341,
739-751.
Kelly, S.J., Li, J., Setlow, P. and Jedrzejas,
M.J. (2007) Structure, flexibility, and mechanism of the Bacillus
stearothermophilus RecU Holliday junction resolvase. Proteins, 68, 961-971.
Khvorova, A., Lescoute, A., Westhof, E. and
Jayasena, S.D. (2003) Sequence elements outside the hammerhead ribozyme
catalytic core enable intracellular activity. Nature Struct. Biol., 10,
1-5.
Klein, D.J., Been, M.D. and Ferré-D'Amaré, A.R.
(2007) Essential role of an active-site guanine in glmS ribozyme catalysis. J. Amer. Chem. Soc., 129, 14858-14859.
Klein, D.J., Schmeing, T.M., Moore, P.B. and
Steitz, T.A. (2001) The kink-turn: a new RNA secondary structure motif. EMBO J., 20, 4214-4221.
Kuzmin, Y.I., Da Costa, C.P., Cottrell, J.W.
and Fedor, M.J. (2005) Role of an active site adenine in hairpin ribozyme
catalysis. J. Molec. Biol., 349, 989-1010.
Kuzmin, Y.I., Da Costa, C.P. and Fedor, M.J.
(2004) Role of an active site guanine in hairpin ribozyme catalysis probed by
exogenous nucleobase rescue. J. Molec.
Biol., 340, 233-251.
Lafontaine, D.A., Norman, D.G. and Lilley,
D.M.J. (2001a) Structure, folding and activity of the VS ribozyme : Importance
of the 2-3-6 helical junction. EMBO J.,
20, 1415-1424.
Lafontaine, D.A., Norman, D.G. and Lilley,
D.M.J. (2002) The global structure of the VS ribozyme. EMBO J., 21, 2461-2471.
Lafontaine, D.A., Wilson, T.J., Norman, D.G.
and Lilley, D.M.J. (2001b) The A730 loop is an important component of the
active site of the VS ribozyme. J. Molec.
Biol., 312, 663-674.
Lescoute, A. and Westhof, E. (2006) Topology of
three-way junctions in folded RNAs. RNA,
12, 83-93.
Lilley, D.M.J. (2000) Structures of helical
junctions in nucleic acids. Quart. Rev.
Biophys., 33, 109-159.
Lilley, D.M.J. (2008) Analysis of branched
nucleic acid structure using comparative gel electrophoresis. Quart. Rev. Biophys., 41, 1-39.
Lilley, D.M.J. (2009a) Comparative gel
electrophoresis analysis of helical junctions in RNA. Meth. Enzymol., 469,
143-177.
Lilley, D.M.J. (2009b) The structure and
folding of branched RNA analyzed by fluorescence resonance energy transfer. Meth. Enzymol., 469, 159-187.
Lilley, D.M.J., Clegg, R.M., Diekmann, S.,
Seeman, N.C., von Kitzing, E. and Hagerman, P. (1995) Nomenclature Committee of
the International Union of Biochemistry: A nomenclature of junctions and
branchpoints in nucleic acids. Recommendations 1994. Eur. J. Biochem., 230,
1-2.
Lilley, D.M.J. and Eckstein, F. (eds.). (2008) Ribozymes and RNA Catalysis. Royal Soc.
Chemistry, Cambridge.
Lilley, D.M.J. and White, M.F. (2000) Resolving
the relationships of resolving enzymes. Proc.
Natl. Acad. Sci. USA, 97, 9351-9353.
Lilley, D.M.J. and White, M.F. (2001) The
junction-resolving enzymes. Nature Rev.
Molec. Cell Biol., 2, 433-443.
Lipfert, J., Ouellet, J., Norman, D.G.,
Doniach, S. and Lilley, D.M.J. (2008) The complete VS ribozyme in solution
studied by small-angle X-ray scattering. Structure,
16, 1357-1367.
Liu, J., Déclais, A.-C. and Lilley, D.M.J.
(2004) Electrostatic interactions and the folding of the four-way DNA junction:
analysis by selective methyl phosphonate substitution. J. Molec. Biol., 343,
851–864.
Liu, J., Déclais, A.-C., McKinney, S.A., Ha,
T., Norman, D.G. and Lilley, D.M.J. (2005) Stereospecific effects determine the
structure of a four-way DNA junction. Chem.
Biol., 12, 217-228.
Liu, J., Déclais, A.C. and Lilley, D.M.J.
(2006) Mechanistic aspects of the DNA junction-resolving enzyme T7 endonuclease
I. Biochemistry, 45, 3934-3942.
Liu, J. and Lilley, D.M.J. (2007) The role of
specific 2'-hydroxyl groups in the stabilization of the folded conformation of
kink-turn RNA. RNA, 13, 200-210.
Liu, S., Bokinsky, G., Walter, N.G. and Zhuang,
X. (2007) Dissecting the multistep reaction pathway of an RNA enzyme by
single-molecule kinetic "fingerprinting". Proc. Natl. Acad. Sci. USA, 104,
12634-12639.
MacKay, C., Déclais, A.-C., Lundin, C.,
Agostinho, A., Deans, A.J., MacArtney, T.J., Hofmann, K., Gartner, A., West,
S.C., Helleday, T., Lilley, D.M.J. and Rouse, J. (2010) Identification of
KIAA1018/FAN1, a DNA repair nuclease recruited to DNA damage by monoubiquitinated
FANCD2. Cell, 142, 65-76.
Martick, M. and Scott, W.G. (2006) Tertiary
contacts distant from the active site prime a ribozyme for catalysis. Cell, 126, 309-320.
McGregor, N., Ayora, S., Sedelnikova, S.,
Carrasco, B., Alonso, J.C., Thaw, P. and Rafferty, J. (2005) The structure of
Bacillus subtilis RecU Holliday junction resolvase and its role in substrate
selection and sequence-specific cleavage. Structure,
13, 1341-1351.
McKinney, S.A., Déclais, A.-C., Lilley, D.M.J.
and Ha, T. (2003) Structural dynamics of individual Holliday junctions. Nature Struct. Biol., 10, 93-97.
McKinney, S.A., Freeman, A.D., Lilley, D.M.J.
and Ha, T. (2005) Observing spontaneous branch migration of Holliday junctions
one step at a time. Proc. Natl. Acad.
Sci. USA, 102, 5715-5720.
McLeod, A.C. and Lilley, D.M.J. (2004)
Efficient, pH-dependent RNA ligation by the VS ribozyme in trans. Biochemistry, 43, 1118 - 1125.
Melcher, S.E., Wilson , T.J. and Lilley ,
D.M.J. (2003) The dynamic nature of the four-way junction of the hepatitis C
virus IRES. RNA, 9, 809-820.
Middleton, C.L., Parker, J.L., Richard, D.J.,
White, M.F. and Bond, C.S. (2004) Substrate recognition and catalysis by the
Holliday junction resolving enzyme Hje. Nucleic
Acids Res., 32, 5442-5451.
Mřllegaard, N.E., Murchie, A.I.H., Lilley,
D.M.J. and Nielsen, P.E. (1994) Uranyl photoprobing of a four-way DNA junction:
Evidence for specific metal ion binding. EMBO
J., 13, 1508-1513.
Munoz, I.M., Hain, K., Déclais, A.-C.,
Gardiner, M., Toh, G.W., Sanchez-Pulido, L., Heuckmann, J., Toth, R.,
Macartney, T., Eppink, B., Kanaar, R., Ponting, C., Lilley, D.M.J. and Rouse,
J. (2009) Coordination of structure–specific nucleases by human SLX4/BTBD12 is
essential for DNA repair. Molec. Cell,
35, 116-127.
Murchie, A.I.H., Clegg, R.M., von Kitzing, E.,
Duckett, D.R., Diekmann, S. and Lilley, D.M.J. (1989) Fluorescence energy
transfer shows that the four-way DNA junction is a right-handed cross of
antiparallel molecules. Nature, 341, 763-766.
Murchie, A.I.H., Thomson, J.B., Walter, F. and
Lilley, D.M.J. (1998) Folding of the hairpin ribozyme in its natural
conformation achieves close physical proximity of the loops. Molec. Cell, 1, 873-881.
Murray, J.B., Seyhan, A.A., Walter, N.G.,
Burke, J.M. and Scott, W.G. (1998) The hammerhead, hairpin and VS ribozymes are
catalytically proficient in monovalent cations alone. Chem. & Biol., 5,
587-595.
Nahas, M.K., Wilson, T.J., Hohng, S., Jarvie,
K., Lilley, D.M.J. and Ha, T. (2004) Observation of internal cleavage and
ligation reactions of a ribozyme. Nature
Struct. Molec. Biol., 11,
1107-1113.
Nesbitt, S., Hegg, L.A. and Fedor, M.J. (1997)
An unusual pH-independent and metal-ion-independent mechanism for hairpin
ribozyme catalysis. Chem. & Biol.,
4, 619-630.
Newman, M., Lunnen, K., Wilson, G., Greci, J.,
Schildkraut, I. and Phillips, S.E.V. (1998) Crystal structure of restriction
endonuclease BglI bound to its interrupted DNA recognition sequence.
EMBO J., 17, 5466-5476.
Nishino, T., Komori, K., Tsuchiya, D., Ishino,
Y. and Morikawa, K. (2001) Crystal structure of the archaeal Holliday junction
resolvase Hjc and implications for DNA recognition. Structure, 9, 197-204.
Norman, D.G., Grainger, R.J., Uhrin, D. and
Lilley, D.M.J. (2000) The location of Cyanine-3 on double-stranded DNA;
importance for fluorescence resonance energy transfer studies. Biochemistry, 39, 6317-6324.
Nowotny, M., Gaidamakov, S.A., Crouch, R.J. and
Yang, W. (2005) Crystal structures of RNase H bound to an RNA/DNA hybrid:
substrate specificity and metal-dependent catalysis. Cell, 121, 1005-1016.
Ortiz-Lombardía, M., González, A., Erijta, R.,
Aymamí, J., Azorín, F. and Coll, M. (1999) Crystal structure of a DNA Holliday
junction. Nature Struct. Biol., 6, 913-917.
Ouellet, J., Melcher, S.E., Iqbal, A., Ding, Y.
and Lilley, D.M.J. (2010) Structure of the three-way helical junction of the
hepatitis C virus IRES element. RNA, 16, 1597-1609
Ouellet, J., Schorr, S., Iqbal, A., Wilson,
T.J. and Lilley, D.M.J. (2011) Orientation of cyanine fluorophores terminally
attached to DNA via long, flexible tethers. Biophys.
J., 101, 1148-1154.
Pena, V., Rozov, A., Fabrizio, P., Luhrmann, R.
and Wahl, M.C. (2008) Structure and function of an RNase H domain at the heart
of the spliceosome. EMBO J., 27, 2929-2940.
Penedo, J.C., Wilson, T.J., Jayasena, S.D.,
Khvorova, A. and Lilley, D.M.J. (2004) Folding of the natural hammerhead
ribozyme is enhanced by interaction of auxiliary elements. RNA, 10, 880-888.
Pinard, R., Hampel, K.J., Heckman, J.E.,
Lambert, D., Chan, P.A., Major, F. and Burke, J.M. (2001) Functional
involvement of G8 in the hairpin ribozyme cleavage mechanism. EMBO J., 20, 6434-6442.
Pley, H.W., Flaherty, K.M. and McKay, D.B.
(1994) Three-dimensional structure of a hammerhead ribozyme. Nature, 372, 68-74.
Potter, H. and Dressler, D. (1978) In vitro system from Escherichia coli that catalyses
generalised genetic recombination. Proc.
Natl. Acad. Sci. USA, 75,
3698-3702.
Raaijmakers, H., Vix, O., Toro, I., Golz, S.,
Kemper, B. and Suck, D. (1999) X-ray structure of T4 endonuclease VII: a DNA
junction resolvase with a novel fold and unusual domain-swapped dimer
architecture. EMBO J., 18, 1447-1458.
Ritchie, D.B., Schellenberg, M.J., Gesner,
E.M., Raithatha, S.A., Stuart, D.T. and Macmillan, A.M. (2008) Structural
elucidation of a PRP8 core domain from the heart of the spliceosome. Nature Struct. Mol. Biol., 15, 1199-1205.
Rupert, P.B. and Ferré-D'Amaré, A.R. (2001)
Crystal structure of a hairpin ribozyme-inhibitor complex with implications for
catalysis. Nature, 410, 780-786.
Rupert, P.B., Massey, A.P., Sigurdsson, S.T.
and Ferré-D'Amaré, A.R. (2002) Transition state stabilization by a catalytic
RNA. Science, 298, 1421-1424.
Ryder, S.P., Oyelere, A.K., Padilla, J.L.,
Klostermeier, D., Millar, D.P. and Strobel, S.A. (2001) Investigation of
adenosine base ionization in the hairpin ribozyme by nucleotide analog
interference mapping. RNA, 7, 1454-1463.
Sanborn, M.E., Connolly, B.K., Gurunathan, K.
and Levitus, M. (2007) Fluorescence properties and photophysics of the
sulfoindocyanine Cy3 linked covalently to DNA. J. Phys. Chem. B, 111,
11064-11074.
Schroeder, K.T., Daldrop, P. and Lilley, D.M.J.
(2011) RNA tertiary interactions in a riboswitch stabilize the structure of a
kink turn. Structure, 19, 1233-1240.
Schroeder, K.T. and Lilley, D.M.J. (2009)
Ion-induced folding of a kink turn that departs from the conventional sequence.
Nucleic Acids Res., 37, 7281-7289.
Schroeder, K.T., McPhee, S.A., Ouellet, J. and
Lilley, D.M. (2010) A structural database for k-turn motifs in RNA. RNA, 16, 1463-1468.
Scott, W.G., Finch, J.T. and Klug, A. (1995)
The crystal structure of an all-RNA hammerhead ribozyme: A proposed mechanism
for RNA catalytic cleavage. Cell, 81, 991-1002.
Smith, M.D. and Collins, R.A. (2007) Evidence
for proton transfer in the rate-limiting step of a fast-cleaving Varkud
satellite ribozyme. Proc. Natl. Acad.
Sci. USA, 104, 5818-5823.
Sood, V.D., Beattie, T.L. and Collins, R.A. (1998)
Identification of phosphate groups involved in metal binding and tertiary
interactions in the core of the Neurospora
VS ribozyme. J. Molec. Biol., 282, 741-750.
Sood, V.D. and Collins, R.A. (2002)
Identification of the catalytic subdomain of the VS ribozyme and evidence for
remarkable sequence tolerance in the active site loop. J. Molec. Biol., 320,
443-454.
Tan, E., Wilson, T.J., Nahas, M.K., Clegg,
R.M., Lilley, D.M.J. and Ha, T. (2003) A four-way junction accelerates hairpin
ribozyme folding via a discrete intermediate. Proc. Natl. Acad. Sci. USA, 100,
9308-9313.
Thomas, J.M. and Perrin, D.M. (2006) Active
site labeling of G8 in the hairpin ribozyme: implications for structure and
mechanism. J. Amer. Chem. Soc., 128, 16540-16545.
Thorpe, J.H., Gale, B.C., Teixeira, S.C. and
Cardin, C.J. (2003) Conformational and hydration effects of site-selective
sodium, calcium and strontium ion binding to the DNA Holliday junction
structure d(TCGGTACCGA)4. J.
Molec. Biol., 327, 97-109.
Turner, B. and Lilley, D.M. (2008) The
importance of G.A hydrogen bonding in the metal ion- and protein-induced
folding of a kink turn RNA. J. Molec.
Biol., 381, 431-442.
Turner, B., Melcher, S.E., Wilson, T.J.,
Norman, D.G. and Lilley, D.M.J. (2005) Induced fit of RNA on binding the L7Ae
protein to the kink-turn motif. RNA, 11, 1192-1200.
Urnavicius, L., McPhee, S.A., Lilley, D.M.J.
and Norman, D.G. (2012) The structure of sulfoindocarbocyanine 3 terminally
attached to dsDNA via a long, flexible tether. Biophys. J., 102,
561-569
Walter, F., Murchie, A.I.H. and Lilley, D.M.J.
(1998a) The folding of the four-way RNA junction of the hairpin ribozyme. Biochemistry, 37, 17629-17636.
Walter, F., Murchie, A.I.H., Thomson, J.B. and
Lilley, D.M.J. (1998b) Structure and activity of the hairpin ribozyme in its
natural junction conformation; effect of metal ions. Biochemistry, 37,
14195-14203.
Walter, N.G., Burke, J.M. and Millar, D.P.
(1999) Stability of hairpin ribozyme tertiary structure is governed by the
interdomain junction. Nature Struct.
Biol., 6, 544-549.
Walter, N.G., Hampel, K.J., Brown, K.M. and
Burke, J.M. (1998c) Tertiary structure formation in the hairpin ribozyme
monitored by fluorescence resonance energy transfer. EMBO J., 17, 2378-2391.
West, S.C. (2003) Molecular views of
recombination proteins and their control. Nature
Rev. Mol. Cell Biol., 4,
435-445.
White, M.F. and Lilley, D.M.J. (1997) The
resolving enzyme CCE1 of yeast opens the structure of the four-way DNA
junction. J. Molec. Biol., 266, 122-134.
Wilson, T.J., Li, N.-S., Lu, J., Frederiksen,
J.K., Piccirilli, J.A. and Lilley, D.M.J. (2010) Nucleobase-mediated general
acid-base catalysis in the Varkud satellite ribozyme. Proc. Natl. Acad. Sci. USA 107,
11751-11756.
Wilson, T.J. and Lilley, D.M.J. (2009) The
evolution of ribozyme chemistry. Science,
323, 1436-1438.
Wilson, T.J. and Lilley, D.M.J. (2011) Do the
hairpin and VS ribozymes share a common catalytic mechanism based on general
acid-base catalysis ? A critical assessment of available experimental data. RNA, 17, 213-221.
Wilson, T.J., McLeod, A.C. and Lilley, D.M.J.
(2007) A guanine nucleobase important for catalysis by the VS ribozyme EMBO J.,
26, 2489-2500.
Wilson, T.J., Ouellet, J., Zhao, Z.Y.,
Harusawa, S., Araki, L., Kurihara, T. and Lilley, D.M.J. (2006) Nucleobase
catalysis in the hairpin ribozyme. RNA,
12, 980-987.
Wilson, T.J., Zhao, Z.-Y., Maxwell, K.,
Kontogiannis, L. and Lilley, D.M.J. (2001) Importance of specific nucleotides
in the folding of the natural form of the hairpin ribozyme. Biochemistry, 40, 2291-2302.
Young, K.J., Gill, F. and Grasby, J.A. (1997)
Metal ions play a passive role in the hairpin ribozyme catalysed reaction. Nucleic Acids Res., 25, 3760-3766.
Zhao, Z., McLeod, A., Harusawa, S., Araki, L.,
Yamaguchi, M., Kurihara, T. and Lilley, D.M.J. (2005) Nucleobase participation
in ribozyme catalysis. J. Amer. Chem.
Soc., 127, 5026-5027.
Zhuang, X.W., Kim, H.D., Pereira, M.J.B.,
Babcock, H.P., Walter, N.G. and Chu, S. (2002) Correlating structural dynamics
and function in single ribozyme molecules. Science,
296, 1473-1476.